Interaction between Saikosaponin D, Paeoniflorin, and Human Serum Albumin
Abstract
:1. Introduction
2. Results and Discussion
2.1. SSD- and PF-Induced Conformational Changes in HSA
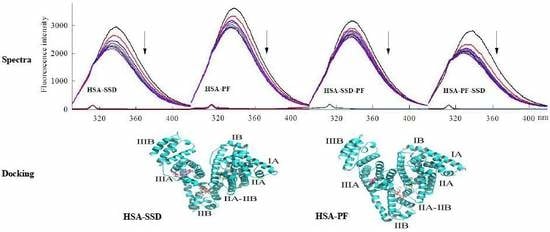
2.1.1. Fluorescence Spectra
2.1.2. Circular Dichroism Studies
2.2. Fluorescence Quenching of HSA by SSD and PF
2.3. Static Quenching of HSA Induced by SSD and PF
2.4. Binding Constants, Sites and Forces
2.4.1. Binding Constants and Numbers of Binding Sites
2.4.2. Identification of SSD or PF Binding Sites
2.4.3. Binding Forces between HSA and SSD/PF
2.5. Energy Transfer Resulting from HSA-SSD and HSA-PF Interaction
2.6. Molecular Docking Studies
3. Materials and Methods
3.1. Materials
3.2. Absorbance Measurements
3.3. Fluorescence Spectra
3.4. Correction of the Internal Filter
3.5. CD Spectra
3.6. Molecular Docking
4. Conclusions
Acknowledgments
Author Contributions
Sample Availability: Not Available. |










| Type | Binding | Detection | Shift and Its Range (nm) | Quenching (%) |
|---|---|---|---|---|
| Binary | HSA-SSD | λex = 280 nm | Left shift 5, 338–333 | 26.55 |
| Δλ = 15 nm | No shift | 7.43 | ||
| Δλ = 60 nm | Left shift 2, 279–277 | 24.34 | ||
| HSA-PF | λex = 280 nm | Left shift 3, 337–334 | 19.42 | |
| Δλ = 15 nm | No shift | 8.29 | ||
| Δλ = 60 nm | Left shift 2, 279–277 | 17.66 | ||
| Terniary | HSA-PF-SSD | λex = 280 nm | Left shift 13, 339–326 | 28.63 |
| HSA-SSD-PF | λex = 280 nm | Left shift 3, 336–333 | 18.19 | |
| Tyrosine | Tyr-SSD | Δλ = 15 nm | No shift | 2.82 |
| Tyr-PF | Δλ = 15 nm | No shift | 4.31 | |
| Tryptophan | Trp-SSD | Δλ = 60 nm | No shift | 12.82 |
| Trp-PF | Δλ = 60 nm | No shift | 10.38 |
| Binding | Ratio | α-Helix | β-Sheet | β-Turn | Random Coil |
|---|---|---|---|---|---|
| HSA | 52.7 | 7.3 | 10.9 | 29.3 | |
| HSA-SSD | 1:1 | 46.3 | 9.0 | 16.9 | 27.8 |
| HSA-PF | 1:1 | 55.6 | 6.5 | 9.1 | 25.2 |
| HSA-PF-SSD | 1:1:1 | 66.4 | 5.0 | 8.6 | 20.1 |
| Binding | Fluorescence | T (°C) | Detection | Kq (L·mol−1·s−1) | R |
|---|---|---|---|---|---|
| HSA-SSD | Conventional | 20 | λex = 280 nm | 1.6 × 1015 | 0.998 |
| 30 | λex = 280 nm | 1.1 × 1015 | 0.996 | ||
| Synchronous | 20 | Δλ = 15 nm | 4.3 × 1014 | 0.991 | |
| 20 | Δλ = 60 nm | 1.3 × 1015 | 0.994 | ||
| 30 | Δλ = 15 nm | 3.5 × 1014 | 0.993 | ||
| 30 | Δλ = 60 nm | 1.1 × 1015 | 0.991 | ||
| HSA-PF | Conventional | 20 | λex = 280 nm | 1.3 × 1015 | 0.995 |
| 30 | λex = 280 nm | 1.1 × 1015 | 0.993 | ||
| Synchronous | 20 | Δλ = 15 nm | 7.5 × 1013 | 0.992 | |
| 20 | Δλ = 60 nm | 7.0 × 1013 | 0.992 | ||
| 30 | Δλ = 15 nm | 3.2 × 1014 | 0.994 | ||
| 30 | Δλ = 60 nm | 5.8 × 1014 | 0.995 | ||
| HSA-PF-SSD | Conventional | 20 | λex = 280 nm | 8.5 × 1014 | 0.992 |
| 30 | λex = 280 nm | 7.3 × 1014 | 0.994 | ||
| HSA-SSD-PF | Conventional | 20 | λex = 280 nm | 5.9 × 1014 | 0.992 |
| 30 | λex = 280 nm | 6.8 × 1014 | 0.993 |
| Binding | T (°C) | Ka (mol·L−1) ± SD | n | R |
|---|---|---|---|---|
| HSA-SSD | 20 | 6.54 × 103 ± 0.146 | 0.525 | 0.998 |
| 30 | 8.28 × 102 ± 0.015 | 0.464 | 0.998 | |
| HSA-PF-SSD | 20 | 2.46 × 103 ± 0.138 | 0.497 | 0.996 |
| 30 | 0.95 × 102 ± 0.121 | 0.230 | 0.996 | |
| HSA-PF | 20 | 7.39 × 104 ± 0.174 | 0.736 | 0.998 |
| 30 | 4.43 × 103 ± 0.019 | 0.482 | 0.995 | |
| HSA-SSD-PF | 20 | 9.58 × 102 ± 0.024 | 0.324 | 0.992 |
| 30 | 3.24 × 102 ± 0.031 | 0.263 | 0.991 |
| Drug | °C | Ka | n | ΔG | ΔS | ΔH | Reaction System | Reference |
|---|---|---|---|---|---|---|---|---|
| PF | 25 | 3.56 × 104 | 0.62 | −21.0 | 103.5 | 4.1 | 10–70 μM PF, 10 μM BSA, pH7.4 Tris-HCL | [34] |
| 37 | 3.78 × 104 | 0.61 | −22.0 | |||||
| PF | 15 | 1.91 × 103 | 0.91 | −18.1 | 28.2 | −10.0 | 2.5–12.5 μM PF, 10 μM BHSA, pH7.4 Tris-HCL | [24] |
| 37 | 1.42 × 103 | 0.89 | −18.7 | |||||
| Paeonol | 10 | 1.58 × 104 | 1.01 | −22.7 | 88.7 | 2.4 | 10–70 μM paenol, 1 μM BHSA, pH7.4 Tris-HCL | [35] |
| 40 | 9.68 × 102 | 0.84 | −17.9 | 64.9 | ||||
| Paeonol | 4 | 1.33 × 104 | - | −22.0 | −6.3 | −23.8 | pH7.4 PBS | [28] |
| 37 | 3.55 × 103 | - | −21.8 | |||||
| SSC | 26 | 3.72 × 103 | 0.79 | −20.34 | −90.3 | −47.3 | 1–15 μM PF, 2 μM BHSA, pH7.4 PBS | [26] |
| Fixed | Binding | Shift and Range (nm) | Quenching (%) | Ka (mol·L−1) ± SD | n | R |
|---|---|---|---|---|---|---|
| Probe | HSA-SSD | Left shift 5, 338–333 | 26.55 | 6.54 × 103 ± 0.146 | 0.525 | 0.998 |
| HSA-War-SSD | Left 5, 337–332 | 29.93 | 3.18 ± 0.016 | 0.224 | 0.995 | |
| HSA-Ibu-SSD | Left 5, 338–333 | 26.12 | 8.42 ± 0.025 | 0.177 | 0.992 | |
| HSA-PF | Left shift 3, 337–334 | 19.42 | 7.39 × 104 ± 0.174 | 0.736 | 0.998 | |
| HSA-War-PF | Left 3, 338–335 | 18.57 | 3.45 ± 0.036 | 0.277 | 0.997 | |
| HSA-Ibu-PF | Left 2, 336–334 | 16.09 | 7.86 ± 0.039 | 0.382 | 0.994 | |
| SSD/PF | HSA-War | Left 3, 338–335 | 19.07 | 2.54 × 102 ± 0.899 | 0.389 | 0.996 |
| HSA-PF-War | Left 4, 340–336 | 14.78 | 4.11 × 105 ± 3.420 | 0.716 | 0.997 | |
| HSA-SSD-War | Left 4, 339–335 | 17.00 | 9.72 × 102 ± 0.448 | 0.469 | 0.996 | |
| HSA-Ibu | Left 3, 338–335 | 17.07 | 4.10 × 102 ± 0.169 | 0.448 | 0.992 | |
| HSA-SSD-Ibu | Left 1, 335–334 | 17.08 | 7.21 × 103 ± 0.368 | 0.520 | 0.994 | |
| HSA-PF-Ibu | Left 3, 338–335 | 19.36 | 4.2 × 105 ± 0.034 | 0.638 | 0.992 |
| Binding | T (°C) | ΔG (kJ·mol−1) | ΔS (J·mol−1·K−1) | ΔH (kJ·mol−1) | J (cm3·L·mol−1) | R0 (nm) | E (J) | r (nm) |
|---|---|---|---|---|---|---|---|---|
| HSA-SSD | 20 | −19.33 | −314.92 | −111.60 | 1.03 × 10−14 | 2.50 | 0.24 | 3.00 |
| 23 | −17.44 | 1.03 × 10−14 | 2.47 | 0.21 | 3.07 | |||
| 27 | −16.45 | 1.04 × 10−14 | 2.06 | 0.13 | 2.83 | |||
| 30 | −16.18 | 1.04 × 10−14 | 2.47 | 0.27 | 2.92 | |||
| HSA-PF | 20 | −27.64 | −1201.16 | −379.58 | 1.88 × 10−15 | 1.86 | 0.05 | 2.27 |
| 23 | −22.19 | 1.91 × 10−15 | 2.02 | 0.08 | 3.02 | |||
| 27 | −21.29 | 1.91 × 10−15 | 2.05 | 0.12 | 2.85 | |||
| 30 | −15.63 | 1.92 × 10−15 | 1.86 | 0.23 | 2.27 |
| Site | Amino Acid Residue | Atoms * | Bond Length | Trp or Tyr around the Pocket | |
|---|---|---|---|---|---|
| SSD | Site I | Lys199 | CH2OH (7) | 2.9 Å | Tyr150 Tyr452 |
| Trp214 | CH2CH2OH (24) | 3.4 Å | |||
| Arg218 | CH2OH (35) | 2.5 Å | |||
| Glu292 | CH2OH (36) | 2.6 Å | |||
| Site II | Asn391 | CH2CH2OH (24) | 2.3 Å | Trp214 | |
| Arg410 | CH2OH (34) | 3.1 Å | |||
| IIA-IIB | Arg209 | CH2CH2OH (37) | 3.3 Å | ||
| PF | Site I | Lys199 | O (21) | 2.9 Å | Tyr150 Trp214 |
| Lys199 | O (27) | 3.0 Å | |||
| Arg257 | O (14) | 3.4 Å | |||
| Arg218 | CH2OH (25) | 2.3 Å | |||
| Ala291 | CH2OH (23) | 2.4 Å | |||
| Glu292 | O (10) | 2.9 Å | |||
| Site II | Asn391 | O (14) | 3.2 Å | Trp214 | |
| Asn391 | CH2OH (23) | 2.5 Å | |||
| Lys414 | O (27) | 3.5 Å | |||
| IIA-IIB | Arg209 | CH2OH (24) | 2.2 Å | Tyr411 | |
| Glu354 | CH2CH2OH (26) | 2.0 Å |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liang, G.-W.; Chen, Y.-C.; Wang, Y.; Wang, H.-M.; Pan, X.-Y.; Chen, P.-H.; Niu, Q.-X. Interaction between Saikosaponin D, Paeoniflorin, and Human Serum Albumin. Molecules 2018, 23, 249. https://doi.org/10.3390/molecules23020249
Liang G-W, Chen Y-C, Wang Y, Wang H-M, Pan X-Y, Chen P-H, Niu Q-X. Interaction between Saikosaponin D, Paeoniflorin, and Human Serum Albumin. Molecules. 2018; 23(2):249. https://doi.org/10.3390/molecules23020249
Chicago/Turabian StyleLiang, Guo-Wu, Yi-Cun Chen, Yi Wang, Hong-Mei Wang, **ang-Yu Pan, Pei-Hong Chen, and Qing-**a Niu. 2018. "Interaction between Saikosaponin D, Paeoniflorin, and Human Serum Albumin" Molecules 23, no. 2: 249. https://doi.org/10.3390/molecules23020249