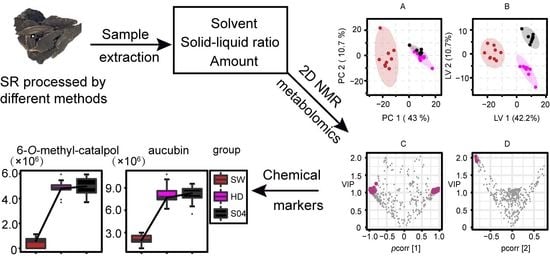
Effect of Different Processing Methods on the Chemical Constituents of Scrophulariae Radix as Revealed by 2D NMR-Based Metabolomics
Abstract
:1. Introduction
2. Results and Discussion
2.1. Samples Collection and Processing
2.2. Optimization of Extraction Conditions
2.2.1. Selecting the Optimum Extraction Solvent
2.2.2. Optimization of the Solid/Liquid Ratio of the Extraction
2.2.3. Optimum Sample Amount Used per NMR Sample
2.3. Comparison of SR Being Steamed Processing for Different Time
2.3.1. Multivariate Statistical Analysis
2.3.2. Change of Chemical Constituents of SR Steamed for Different Durations
| Feature | δH | δC | Assignment | Feature | δH | δC | Assignment | Feature | δH | δC | Assignment |
|---|---|---|---|---|---|---|---|---|---|---|---|
| F115 | 2.72 | 46.66 | aucubin | F121 | 3.02 | 49.38 | aucubin | F124 | 5.05 | 99.01 | aucubin |
| F128 | 5.81 | 131.75 | aucubin | F127 | 5.12 | 107.85 | aucubin | F135 | 6.32 | 142.93 | aucubin |
| F179 | 4.33 | 62.72 | aucubin | F196 | 4.35 | 62.72 | aucubin | F283 | 3.84 | 79.19 | harpagoside |
| F299 | 5.01 | 108.14 | harpagoside | F346 | 2.95 | 56.61 | harpagoside | F372 | 6.16 | 96.43 | harpagoside |
| F408 | 6.47 | 145.71 | harpagoside | F146 | 1.52 | 24.36 | harpagoside | F558 | 2.29 | 47.69 | harpagoside |
| F581 | 2.04 | 47.68 | harpagoside | F773 | 6.55 | 121.43 | harpagoside, cinnamic acid | F225 | 7.63 | 131.07 | harpagoside, cinnamic acid |
| F679 | 7.69 | 148.31 | harpagoside, cinnamic acid | F101 | 4.72 | 101.44 | aucubin, harpagoside | F118 | 1.26 | 26.82 | harpagide |
| F242 | 5.03 | 109.70 | harpagide | F216 | 3.78 | 79.47 | harpagide | F219 | 4.67 | 101.09 | harpagide |
| F279 | 6.37 | 144.22 | harpagide | F254 | 2.57 | 60.43 | harpagide | F273 | 5.73 | 95.26 | harpagide |
| F375 | 1.95 | 48.66 | harpagide | F294 | 1.85 | 48.66 | harpagide | F607 | 1.83 | 48.66 | harpagide |
| F495 | 1.97 | 48.66 | harpagide | F106 | 3.30 | 76.01 | aucubin, harpagoside, harpagide | F268 | 3.31 | 75.64 | aucubin, harpagoside, harpagide |
| F244 | 3.35 | 72.89 | aucubin, harpagoside, harpagide, glucose | F220 | 3.77 | 61.50 | 6-O-methyl-catalpol | F218 | 2.35 | 38.86 | 6-O-methyl-catalpol |
| F229 | 6.40 | 143.71 | 6-O-methyl-catalpol | F207 | 5.08 | 105.85 | 6-O-methyl-catalpol | F520 | 4.18 | 63.00 | 6-O-methyl-catalpol |
| F138 | 4.82 | 101.35 | 6-O-methyl-catalpol | F201 | 5.05 | 97.23 | 6-O-methyl-catalpol | F123 | 4.20 | 62.79 | 6-O-methyl-catalpol |
| F184 | 3.35 | 76.10 | 6-O-methyl-catalpol | F194 | 2.60 | 44.63 | 6-O-methyl-catalpol | F178 | 4.23 | 62.72 | 6-O-methyl-catalpol |
| F111 | 3.48 | 78.90 | aucubin, harpagoside, harpagide, 6-O-methyl-catalpol, glucuronic acid | F107 | 3.38 | 79.20 | aucubin, harpagoside, harpagide, 6-O-methyl-catalpol, glucose | F108 | 3.37 | 79.39 | aucubin, harpagoside, harpagide, 6-O-methyl-catalpol |
| F42 | 3.45 | 79.22 | aucubin, harpagoside, harpagide, 6-O-methyl-catalpol, glucuronic acid | F300 | 3.41 | 79.25 | aucubin, harpagoside, harpagide, 6-O-methyl-catalpol | F228 | 3.41 | 79.45 | aucubin, harpagoside, harpagide, 6-O-methyl-catalpol, glucose |
| F649 | 1.53 | 27.47 | isoleucine | F367 | 3.39 | 78.91 | aucubin, harpagoside, harpagide, 6-O-methyl-catalpol, glucose | F460 | 3.61 | 62.61 | isoleucine |
| F238 | 0.96 | 14.06 | isoleucine | F611 | 1.98 | 39.00 | isoleucine | F224 | 1.03 | 17.53 | isoleucine |
| F159 | 7.18 | 133.57 | tyrosine | F330 | 2.65 | 32.01 | methionine | F1423 | 3.00 | 38.63 | tyrosine |
| F659 | 3.20 | 38.64 | tyrosine | F147 | 6.85 | 118.65 | tyrosine | F248 | 3.88 | 59.18 | tyrosine |
| F339 | 2.05 | 28.38 | glutamine | F357 | 3.01 | 38.65 | tyrosine | F389 | 3.22 | 38.65 | tyrosine |
| F186 | 3.53 | 63.50 | threonine, valine | F261 | 2.09 | 28.39 | glutamine | F145 | 2.07 | 28.41 | glutamine |
| F596 | 1.76 | 43.04 | leucine | F95 | 1.35 | 22.64 | threonine | F235 | 0.99 | 25.00 | leucine |
| F297 | 1.73 | 27.11 | leucine, arginine | F230 | 0.97 | 23.83 | leucine | F637 | 1.66 | 43.03 | leucine |
| F198 | 1.94 | 30.71 | arginine | F350 | 1.67 | 27.09 | arginine | F470 | 3.21 | 43.69 | arginine |
| F40 | 3.83 | 75.61 | sucrose | F213 | 1.90 | 30.67 | arginine | F622 | 3.82 | 62.76 | sucrose |
| F31 | 3.80 | 63.37 | sucrose | F53 | 3.44 | 72.46 | sucrose | F28 | 3.83 | 65.34 | sucrose |
| F19 | 3.76 | 75.90 | sucrose | F55 | 3.78 | 63.39 | sucrose | F18 | 3.54 | 74.19 | sucrose |
| F5 | 3.81 | 65.32 | sucrose, stachyose, raffinose | F2 | 3.66 | 64.87 | sucrose, stachyose | F12 | 3.78 | 65.34 | sucrose, stachyose, raffinose |
| F11 | 5.43 | 95.10 | sucrose, stachyose, raffinose | F6 | 4.18 | 79.84 | sucrose, stachyose, raffinose | F9 | 3.85 | 84.66 | sucrose, stachyose, raffinose |
| F23 | 4.95 | 101.65 | stachyose, raffinose | F65 | 5.41 | 95.23 | sucrose, stachyose, raffinose | F20 | 4.12 | 71.75 | stachyose |
| F25 | 3.67 | 69.01 | stachyose, raffinose | F22 | 4.02 | 69.01 | stachyose, raffinose | F21 | 4.06 | 74.43 | stachyose, raffinose |
| F46 | 3.23 | 77.36 | glucose, glucuronic acid | F56 | 3.71 | 76.20 | sucrose, glucose, glucuronic acid | F51 | 5.21 | 95.42 | glucuronic acid |
| F258 | 3.25 | 77.38 | glucose | F116 | 3.81 | 74.47 | glucose | F54 | 3.48 | 74.80 | glucose |
| F137 | 3.92 | 63.97 | glucose | F103 | 4.58 | 99.25 | β-glucose | F102 | 3.21 | 77.44 | glucose |
| F97 | 4.01 | 66.26 | fructose | F155 | 3.46 | 74.92 | glucose | F57 | 4.03 | 66.26 | fructose |
| F328 | 3.70 | 67.08 | fructose | F44 | 3.81 | 70.65 | fructose | F180 | 3.98 | 79.17 | fructose |
| F41 | 3.71 | 67.08 | fructose | F74 | 3.70 | 66.25 | fructose | F154 | 4.07 | 85.43 | fructose |
| F61 | 3.55 | 65.84 | fructose | F35 | 3.53 | 67.08 | fructose | F321 | 3.51 | 67.08 | fructose |
| F48 | 3.68 | 66.24 | fructose | F68 | 4.08 | 78.69 | fructose | F163 | 3.79 | 70.65 | fructose |
| F71 | 3.53 | 65.82 | fructose | F692 | 3.02 | 41.93 | 4-aminobutyric acid | F199 | 2.32 | 37.41 | 4-aminobutyric acid |
| F175 | 3.01 | 42.37 | 4-aminobutyric acid | F192 | 1.91 | 26.53 | 4-aminobutyric acid | F624 | 2.29 | 32.19 | valine |
| F237 | 1.06 | 20.98 | valine | F232 | 1.02 | 19.63 | valine | F223 | 3.92 | 79.83 | |
| F260 | 3.24 | 56.82 | F386 | 1.94 | 25.75 | F182 | 3.21 | 56.76 | |||
| F546 | 4.05 | 58.63 | F758 | 2.67 | 39.47 | F92 | 4.14 | 61.30 | |||
| F34 | 4.62 | 99.40 | F946 | 2.83 | 39.45 | F975 | 3.12 | 44.34 | |||
| F791 | 3.81 | 60.65 | F52 | 3.73 | 68.65 | F503 | 3.51 | 70.54 | |||
| F473 | 3.86 | 55.36 | F740 | 6.78 | 118.64 | F864 | 2.81 | 39.47 | |||
| F292 | 2.50 | 28.39 | F156 | 2.48 | 28.41 | F309 | 2.41 | 36.86 | |||
| F872 | 6.67 | 123.63 | F176 | 2.40 | 32.70 | F853 | 6.79 | 119.26 | |||
| F90 | 2.38 | 32.74 | F181 | 4.02 | 84.29 | F204 | 2.36 | 32.69 | |||
| F285 | 2.46 | 28.39 | F278 | 5.04 | 110.79 | F642 | 3.59 | 49.68 | |||
| F708 | 2.11 | 32.99 | F288 | 3.59 | 72.65 | F49 | 3.98 | 68.66 | |||
| F36 | 3.85 | 72.91 | F315 | 5.11 | 110.38 | F120 | 3.50 | 74.41 | |||
| F487 | 1.30 | 19.68 | F322 | 4.00 | 86.92 | F500 | 3.67 | 56.35 | |||
| F502 | 4.18 | 85.47 | F354 | 4.24 | 82.63 | F545 | 2.82 | 37.70 | |||
| F505 | 3.82 | 55.44 | F480 | 5.06 | 110.70 | F772 | 3.40 | 58.90 | |||
| F636 | 2.04 | 24.55 | F557 | 4.25 | 84.47 | F1400 | 5.43 | 75.92 | |||
| F405 | 3.25 | 77.58 | F608 | 3.96 | 80.14 | F1430 | 4.97 | 72.81 | |||
| F699 | 2.21 | 32.98 | F616 | 4.10 | 84.97 | F1483 | 5.43 | 74.44 | |||
| F447 | 2.75 | 39.54 | F645 | 4.09 | 84.02 | F117 | 3.51 | 60.15 | |||
| F43 | 3.59 | 77.32 | F646 | 3.91 | 76.55 | F188 | 3.76 | 74.20 | |||
| F723 | 3.42 | 57.74 | F777 | 1.79 | 29.00 | F195 | 3.75 | 89.84 | |||
| F66 | 3.80 | 84.11 | F921 | 2.15 | 23.05 | F739 | 6.56 | 138.61 | |||
| F72 | 4.00 | 73.00 | F745 | 1.91 | 33.03 | F287 | 4.18 | 77.06 | |||
| F89 | 4.08 | 77.78 | F1209 | 5.90 | 128.65 | F457 | 4.06 | 84.65 | |||
| F349 | 2.14 | 16.84 | F13 | 4.06 | 77.06 | F29 | 3.83 | 71.78 | |||
| F32 | 4.16 | 71.74 | F125 | 4.48 | 84.00 | F206 | 4.06 | 79.80 | |||
| F174 | 3.87 | 73.28 | F422 | 3.56 | 63.35 | F750 | 5.43 | 85.29 | |||
| F193 | 3.61 | 66.25 | F845 | 4.29 | 84.69 | F684 | 3.36 | 49.67 | |||
| F256 | 4.10 | 84.45 | F932 | 4.16 | 85.44 |
2.4. Comparison of SR Processed by SW, HD, and S04
2.4.1. Multivariate Statistical Analysis
2.4.2. Identification of Differential Metabolites of SW, HD, and S04
| Feature | δH | δC | Assignment | Feature | δH | δC | Assignment | Feature | δH | δC | Assignment |
|---|---|---|---|---|---|---|---|---|---|---|---|
| F115 | 2.72 | 46.66 | aucubin | F121 | 3.02 | 49.38 | aucubin | F124 | 5.05 | 99.01 | aucubin |
| F135 | 6.32 | 142.93 | aucubin | F128 | 5.81 | 131.75 | aucubin | F179 | 4.33 | 62.72 | aucubin |
| F127 | 5.12 | 107.85 | aucubin | F685 | 2.27 | 47.68 | harpagoside | F299 | 5.01 | 108.14 | harpagoside |
| F408 | 6.47 | 145.71 | harpagoside | F372 | 6.16 | 96.43 | harpagoside | F346 | 2.95 | 56.61 | harpagoside |
| F146 | 1.52 | 24.36 | harpagoside | F625 | 2.06 | 47.68 | harpagoside | F283 | 3.84 | 79.19 | harpagoside |
| F712 | 7.66 | 148.31 | harpagoside, cinnamic acid | F225 | 7.63 | 131.07 | harpagoside, cinnamic acid | F773 | 6.55 | 121.43 | harpagoside, cinnamic acid |
| F671 | 6.53 | 121.43 | harpagoside, cinnamic acid | F101 | 4.72 | 101.44 | aucubin, harpagoside | F244 | 3.35 | 72.89 | aucubin, harpagoside, glucose |
| F138 | 4.82 | 101.35 | 6-O-methyl-catalpol | F207 | 5.08 | 105.85 | 6-O-methyl-catalpol | F194 | 2.60 | 44.63 | 6-O-methyl-catalpol |
| F123 | 4.20 | 62.79 | 6-O-methyl-catalpol | F184 | 3.35 | 76.10 | 6-O-methyl-catalpol | F229 | 6.40 | 143.71 | 6-O-methyl-catalpol |
| F201 | 5.05 | 97.23 | 6-O-methyl-catalpol | F218 | 2.35 | 38.86 | 6-O-methyl-catalpol | F220 | 3.77 | 61.50 | 6-O-methyl-catalpol |
| F42 | 3.45 | 79.22 | aucubin, harpagoside, 6-O-methyl-catalpol, glucose | F333 | 3.39 | 79.72 | aucubin, harpagoside, 6-O-methyl-catalpol, glucose | F111 | 3.48 | 78.90 | aucubin, harpagoside, 6-O-methyl-catalpol |
| F108 | 3.37 | 79.39 | aucubin, harpagoside, 6-O-methyl-catalpol | F367 | 3.39 | 78.91 | aucubin, harpagoside, 6-O-methyl-catalpol, glucose | F130 | 3.46 | 79.14 | aucubin, harpagoside, 6-O-methyl-catalpol |
| F107 | 3.38 | 79.20 | aucubin, harpagoside, 6-O-methyl-catalpol, glucose | F211 | 3.44 | 79.30 | aucubin, harpagoside, 6-O-methyl-catalpol | F494 | 3.80 | 56.89 | methionine |
| F699 | 2.21 | 32.98 | methionine | F349 | 2.14 | 16.84 | methionine | F330 | 2.65 | 32.01 | methionine |
| F708 | 2.11 | 32.99 | methionine | F659 | 3.20 | 38.64 | tyrosine | F1423 | 3.00 | 38.63 | tyrosine |
| F248 | 3.88 | 59.18 | tyrosine | F103 | 4.58 | 99.25 | β-glucose | F150 | 5.19 | 95.24 | α-glucose |
| F102 | 3.21 | 77.44 | glucose | F50 | 3.73 | 57.21 | F1209 | 5.90 | 128.65 | ||
| F88 | 3.98 | 72.38 | F667 | 5.14 | 98.33 | F125 | 4.48 | 84.00 | |||
| F77 | 3.91 | 74.04 | F1055 | 1.58 | 30.33 | F340 | 1.24 | 17.22 | |||
| F117 | 3.51 | 60.15 | F195 | 3.75 | 89.84 | F514 | 1.94 | 24.72 | |||
| F622 | 3.82 | 62.76 | F47 | 1.35 | 23.18 | F610 | 2.36 | 37.79 | |||
| F723 | 3.42 | 57.74 | F606 | 4.01 | 82.57 | F631 | 4.45 | 105.80 | |||
| F988 | 1.93 | 29.66 | F689 | 7.38 | 132.36 | F741 | 3.13 | 51.58 | |||
| F717 | 1.95 | 29.64 | F727 | 1.99 | 29.63 | F760 | 3.35 | 76.83 | |||
| F761 | 3.26 | 55.60 | F772 | 3.40 | 58.90 | F989 | 4.81 | 101.63 | |||
| F777 | 1.79 | 29.00 | F794 | 3.35 | 55.66 | F800 | 4.02 | 66.89 | |||
| F182 | 3.21 | 56.76 | F503 | 3.51 | 70.54 | F546 | 4.05 | 58.63 |
3. Materials and Methods
3.1. Chemicals and Reagents
3.2. Samples Collection and Processing
3.2.1. Fresh SR Raw Material Collection
3.2.2. SR Raw Material Samples Processing
3.3. Optimization of Extraction Conditions
3.4. NMR Sample Preparation and Data Acquisition
3.5. NMR Spectrum Processing and Data Preprocessing
3.6. Chemometric Analysis
3.7. Metabolite Identification and Verification
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Sample Availability
References
- ** laboratory Escherichia coli Strains. Chem. Biodivers. 2013, 10, 1816–1827. [Google Scholar] [CrossRef]
- Ai, Z.P.; Zhang, Y.; Li, X.Y.; Sun, W.L.; Liu, Y.H. Widely targeted metabolomics analysis to reveal transformation mechanism of cistanche deserticola active compounds during steaming and drying processes. Front. Nutr. 2021, 8, 742511. [Google Scholar] [CrossRef]
- **, J.; Lao, J.; Zhou, R.R.; He, W.; Qin, Y.; Zhong, C.; **e, J.; Liu, H.; Wan, D.; Zhang, S.H.; et al. Simultaneous identification and dynamic analysis of saccharides during steam processing of rhizomes of polygonatum cyrtonema by HPLC-QTOF-MS/MS. Molecules 2018, 23, 2855. [Google Scholar] [CrossRef] [Green Version]
- Chylla, R.A.; Hu, K.F.; Effinger, J.J.; Markley, J.L. Deconvolution of two-dimensional NMR spectra by fast maximum likelihood reconstruction: Application to quantitative metabolomics. Anal. Chem. 2011, 83, 4871–4880. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.S.; Kim, S.Y.; Chun, Y.S.; Chun, Y.J.; Shin, S.Y.; Choi, C.H.; Choi, H.K. Characteristics of fecal metabolic profiles in patients with irritable bowel syndrome with predominant diarrhea investigated using H-1-NMR coupled with multivariate statistical analysis. Neurogastroenterol. Motil. 2020, 32, e13830. [Google Scholar] [CrossRef] [PubMed]
- Brahmi, F.; Nguyen, A.T.; Nacoulma, A.P.; Sheridan, H.; Wang, J.F.; Guendouze, N.; Madani, K.; Duez, P. Discrimination of Mentha species grown in different geographical areas of algeria using H-1-NMR-based metabolomics. J. Pharm. Biomed. Anal. 2020, 189, 113430. [Google Scholar] [CrossRef] [PubMed]
- Yan, Y.; Shen, C.X.; Pei, X.P.; Li, A.P.; Du, H.Z.; Qin, X.M.; Du, C.H. Compare the chemical profiling of Ziziphi Spinosae Semen and Ziziphi Mauritianae Semen using untargeted metabolomics. J. Food Compos. Anal. 2020, 94, 103635. [Google Scholar] [CrossRef]
- Gogna, N.; Hamid, N.; Dorai, K. Metabolomic profiling of the phytomedicinal constituents of Carica papaya L. leaves and seeds by H-1 NMR spectroscopy and multivariate statistical analysis. J. Pharm. Biomed. Anal. 2015, 115, 74–85. [Google Scholar] [CrossRef]
- Masetti, O.; Nisini, L.; Ciampa, A.; Dell’Abate, M.T. H-1 NMR spectroscopy coupled with multivariate analysis was applied to investigate Italian cherry tomatoes metabolic profile. J. Chemometr. 2020, 34, e3191. [Google Scholar] [CrossRef]
- Villa-Ruano, N.; Rosas-Bautista, A.; Rico-Arzate, E.; Cruz-Narvaez, Y.; Zepeda-Vallejo, L.G.; Lalaleo, L.; Hidalgo-Martinez, D.; Becerra-Martinez, E. Study of nutritional quality of pomegranate (Punica granatum L.) juice using H-1 NMR-based metabolomic approach: A comparison between conventionally and organically grown fruits. LWT 2020, 134, 110222. [Google Scholar] [CrossRef]
- Huang, T.; Chen, P.; Liu, B.; Li, X.; Lv, X.; Hu, K. NPid: An automatic approach to rapid identification of known natural products in the crude extract of crabapple based on 2D H-1-C-13 heteronuclear correlation spectra of the extract mixture. Anal. Chem. 2020, 92, 10996–11006. [Google Scholar] [CrossRef]





Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Duan, X.; Zhang, M.; Du, H.; Gu, X.; Bai, C.; Zhang, L.; Chen, K.; Hu, K.; Li, Y. Effect of Different Processing Methods on the Chemical Constituents of Scrophulariae Radix as Revealed by 2D NMR-Based Metabolomics. Molecules 2022, 27, 4687. https://doi.org/10.3390/molecules27154687
Duan X, Zhang M, Du H, Gu X, Bai C, Zhang L, Chen K, Hu K, Li Y. Effect of Different Processing Methods on the Chemical Constituents of Scrophulariae Radix as Revealed by 2D NMR-Based Metabolomics. Molecules. 2022; 27(15):4687. https://doi.org/10.3390/molecules27154687
Chicago/Turabian StyleDuan, **aohui, Mina Zhang, Huan Du, **u Gu, Caihong Bai, Liuqiang Zhang, Kaixian Chen, Kaifeng Hu, and Yiming Li. 2022. "Effect of Different Processing Methods on the Chemical Constituents of Scrophulariae Radix as Revealed by 2D NMR-Based Metabolomics" Molecules 27, no. 15: 4687. https://doi.org/10.3390/molecules27154687