Identification of miRNAs and Their Targets in Cotton Inoculated with Verticillium dahliae by High-Throughput Sequencing and Degradome Analysis
Abstract
:1. Introduction
2. Results
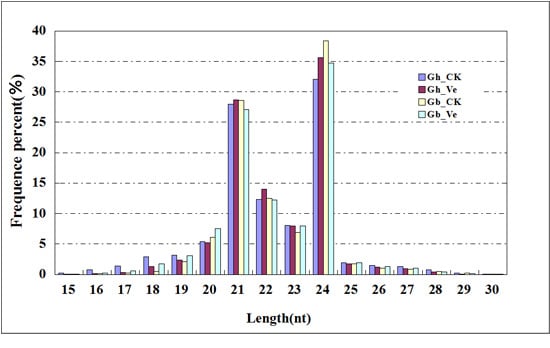
2.1. Overview of Small RNA (sRNA) Library Sequencing
| Category | Library Name | |||
|---|---|---|---|---|
| Gh_CK | Gh_Ve | Gb_CK | Gb_Ve | |
| Total reads | 26,495,780 | 19,511,637 | 20,609,252 | 19,965,211 |
| High quality | 26,409,519 | 19,450,186 | 20,548,220 | 19,903,524 |
| Clean reads | 25,679,650 | 19,296,249 | 20,389,102 | 19,638,276 |
| Unique sRNAs | 7,181,742 | 6,033,991 | 6,729,005 | 6,415,595 |
| Unique sRNA map** to genome | 2,235,867 | 1,855,605 | 2,078,150 | 1,988,015 |
| miRNA | 37,936 | 29,080 | 32,658 | 34,630 |
| Unannotated | 6,264,557 | 5,281,380 | 5,889,075 | 5,583,553 |

2.2. Identification of Known MicroRNAs (miRNAs) by sRNA Sequencing
2.3. Identification of Novel miRNAs in G. hirsutum and G. barbadense
| miRNA | Mature Sequence | LM | Arm | LP | G + C (%) | MFE | Reads Per Million | |||
|---|---|---|---|---|---|---|---|---|---|---|
| Gh_CK | Gh_Ve | Gb_CK | Gb_Ve | |||||||
| novel_miR_1 | TTGACAAGTAAAAGAACATA | 20 | 3p | 147 | 29.25 | −23.44 | 0 | 0.98 | 0.74 | 1.07 |
| novel_miR_2 | CGAAGTCTTGGAAGAGAGTAA | 21 | 3p | 91 | 37.36 | −39.7 | 49.65 | 0 | 24.82 | 30.3 |
| novel_miR_3 | TGGTATTGGAGTGAAGGGAGC | 21 | 5p | 70 | 37.14 | −21.1 | 50.9 | 73.64 | 68.57 | 44.5 |
| novel_miR_4 | TGATTGAGCCGTGCCAATATC | 21 | 3p | 101 | 42.57 | −53.9 | 115.81 | 141.22 | 231.64 | 163.25 |
| novel_miR_5 | GTGGGCGTGCCGGAGTGGTTA | 21 | 5p | 74 | 56.76 | −26.8 | 110.52 | 96.7 | 34.04 | 36.76 |
| novel_miR_6 | CGGACTCTCAAACAGTGGAGGTA | 23 | 5p | 256 | 45.31 | −108.08 | 7.63 | 5.44 | 0 | 12.78 |
| novel_miR_7 | CTGGACTGTCAATGGCCGGCAC | 22 | 3p | 95 | 45.26 | −35.4 | 0 | 0 | 2.45 | 1.63 |
| novel_miR_8 | ATTAGATACTATAGCATGAGACA | 23 | 5p | 289 | 24.57 | −46.7 | 3.66 | 0 | 0 | 9.01 |
| novel_miR_9 | CTGTCGCAGGGGAGATGGCTCGT | 23 | 5p | 141 | 58.16 | −66.1 | 9.27 | 8.24 | 0 | 0.51 |
| novel_miR_10 | TCTAATAGAAGAATGACAAATCA | 23 | 5p | 84 | 20.24 | −22 | 0 | 0.93 | 1.52 | 0.81 |
| novel_miR_11 | TGGGAAATGATGACAGCTTA | 20 | 3p | 193 | 29.53 | −44.9 | 4.24 | 0 | 0 | 1.22 |
| novel_miR_12 | GCAGATGATGATAAGAATAGACA | 23 | 5p | 140 | 40.71 | −33.3 | 2.26 | 2.23 | 1.77 | 0.66 |
| novel_miR_13 | GGGCGCCTCTTACTTGGCAGG | 21 | 5p | 132 | 44.7 | −56.5 | 0.9 | 4.3 | 0 | 7.13 |
| novel_miR_14 | ATTAGATACTATAGCGTGAGACA | 23 | 3p | 226 | 37.17 | −61.1 | 0 | 0 | 0 | 7.69 |
| novel_miR_15 | GCTCACTTCTCTTTCTGTCAGTT | 23 | 3p | 107 | 44.86 | −57.7 | 5.33 | 9.74 | 11.87 | 7.84 |
| novel_miR_16 | TGCCTGGCTCCCTGAATGCCA | 21 | 5p | 104 | 50 | −53.3 | 1.75 | 0.73 | 0.49 | 1.32 |
| novel_miR_17 | AGGTGCAGGTGCAGGCGCAGC | 21 | 3p | 146 | 40.41 | −49.62 | 0 | 0 | 7.9 | 1.53 |
| novel_miR_18 | CGGACCCTCTAACAGTGGAGG | 21 | 5p | 214 | 50.47 | −90.3 | 56.93 | 37.47 | 0 | 38.9 |
| novel_miR_19 | TGAGAAAGTGGAGATGGGTGG | 21 | 3p | 128 | 45.31 | −63.9 | 2.18 | 1.71 | 3.87 | 2.6 |
| novel_miR_20 | CTCTGGTTTGACTCATTTGTA | 21 | 5p | 181 | 33.7 | −66.8 | 3.12 | 0 | 0 | 1.63 |
| novel_miR_21 | CAAATGAGTTAGGCGAGAGGT | 21 | 3p | 170 | 33.53 | −63.56 | 0 | 13.42 | 0 | 7.08 |
| novel_miR_22 | TCAAGCAAATCAAGGAAAGGCC | 22 | 3p | 344 | 38.66 | −113.5 | 0.55 | 0.47 | 1.13 | 0.36 |
| novel_miR_23 | TCTGAAAAGCAATAAAGAACACA | 23 | 3p | 94 | 27.66 | −25.7 | 0 | 0 | 2.55 | 1.78 |
| novel_miR_24 | GTGGAAGTTAGGCTGACTTAGGC | 23 | 5p | 79 | 45.57 | −21.7 | 0 | 0 | 0 | 4.12 |
| novel_miR_25 | AGGCAGTCACCTTGGCTAAC | 20 | 5p | 180 | 45.56 | −64.6 | 0 | 4.98 | 0 | 1.68 |
| novel_miR_26 | TTGGATGGGCGGTGTGTTTACTT | 23 | 3p | 246 | 28.46 | −49.4 | 0 | 0 | 0.64 | 1.93 |
| novel_miR_27 | GGCAAGTTGTCCTCGGCTACA | 21 | 3p | 178 | 39.89 | −59.96 | 0 | 1.87 | 0 | 1.12 |
| novel_miR_28 | TCCATATTTCACTATCTCTTA | 21 | 3p | 105 | 32.38 | −48.6 | 11.25 | 16.01 | 72.54 | 22.66 |
| novel_miR_29 | TTGAACACCGAAGTAAAGCCAT | 22 | 5p | 209 | 41.15 | −82.8 | 0 | 0 | 0 | 1.78 |
| novel_miR_30 | TGCCAAATCAGGGAAGCGAA | 20 | 5p | 148 | 41.22 | −52.32 | 0 | 0 | 43.5 | 0 |
| novel_miR_31 | AGGCTGTGATGACGAGAAATTA | 22 | 3p | 237 | 30.38 | −47.71 | 32.94 | 24.1 | 16.92 | 0 |
| novel_miR_32 | TTTCCAATAGAAGAATGACAAAT | 23 | 5p | 140 | 27.14 | −54.8 | 1.36 | 1.09 | 1.03 | 0 |
| novel_miR_33 | AATCTCCCTCAAACGCTTCCAG | 22 | 5p | 118 | 45.76 | −47.84 | 0 | 0 | 9.66 | 0 |
| novel_miR_34 | TTTGGATTGAAGGGAGCTCTA | 21 | 3p | 203 | 42.36 | −92.45 | 0 | 0 | 141.2 | 0 |
| novel_miR_35 | TAGTGAGGATGGGAAATTTGT | 21 | 5p | 124 | 30.65 | −49.3 | 34.93 | 0 | 18.78 | 14 |
| novel_miR_36 | GAGCTTGGAAGTGCATCCGGC | 21 | 5p | 107 | 45.79 | −54.9 | 0 | 0 | 4.07 | 0 |
| novel_miR_37 | TCGCTTCCCTAATTTGGACGA | 21 | 3p | 148 | 41.22 | −52.32 | 36.33 | 31.09 | 0 | 85.09 |
| novel_miR_38 | TGACTCCTAGTACAACGGCCTC | 22 | 3p | 328 | 32.32 | −123.5 | 0 | 0 | 5.84 | 0 |
| novel_miR_39 | TTGAGCCGTGCCAATATCAATC | 22 | 3p | 108 | 47.22 | −49.81 | 0 | 0 | 226.44 | 0 |
| novel_miR_40 | CAAAGAGTAGAGGTATTGTGC | 21 | 5p | 272 | 41.54 | −112 | 0 | 13.89 | 1.23 | 0 |
| novel_miR_41 | ACAGGTTAGTAGAAATTAAGGTT | 23 | 5p | 115 | 41.74 | −22.6 | 0 | 1.09 | 0 | 0 |
| novel_miR_42 | CAGAATGACCAATTTACTCTTTA | 23 | 3p | 199 | 24.12 | −40.4 | 0 | 4.92 | 0 | 0 |
| novel_miR_43 | ACTCTCTTCCAAAGGCTTCAAG | 22 | 5p | 108 | 35.19 | −46 | 0 | 49.91 | 0 | 0 |
| novel_miR_44 | TGGTGCAGGTCGGGAACTGAT | 21 | 5p | 139 | 47.48 | −57.4 | 0 | 5.86 | 0 | 0 |
| novel_miR_45 | GTTCAATAAAGCTGTGGGAAG | 21 | 3p | 138 | 40.58 | −54.6 | 0 | 100.69 | 0 | 0 |
| novel_miR_46 | CAAAAGCAATGAAGAACTGGCCA | 23 | 5p | 331 | 34.14 | −78.9 | 0 | 1.97 | 0 | 0 |
| novel_miR_47 | ACAGTAGAAATGGATGGAATT | 21 | 3p | 153 | 30.07 | −48 | 0 | 3.99 | 0 | 0 |
| novel_miR_48 | TTGGATGGACGGTGCATTTATCT | 23 | 3p | 295 | 37.29 | −62.4 | 0 | 1.04 | 0 | 0 |
| novel_miR_49 | ATTATTGTTAATGTAGGAGGA | 21 | 5p | 371 | 31 | −61.26 | 0 | 1.92 | 0 | 0 |
| novel_miR_50 | TTGTACTAGGAGTCGGATTGC | 21 | 5p | 344 | 33.72 | −143.2 | 0 | 4.15 | 0 | 1.27 |
| novel_miR_51 | TTAGATCAAAGAGCAAACCGG | 21 | 5p | 210 | 36.67 | −95.5 | 1.32 | 0 | 0 | 0 |
| novel_miR_52 | CGAGACTTGCGGTAGAAACAAA | 22 | 3p | 149 | 40.27 | −35.2 | 6.74 | 0 | 0 | 0 |
| novel_miR_53 | TGGAGGCAGCGGTTCATCGATC | 22 | 5p | 110 | 42.73 | −40.1 | 0 | 0 | 0 | 12.37 |
| novel_miR_54 | GGGACTCTCCGGACTGTTTGGTT | 23 | 3p | 148 | 36.49 | −43.3 | 1.05 | 0 | 0 | 0 |
| novel_miR_55 | TGCCTGGCTCCCTGTATGCCA | 21 | 5p | 103 | 47.57 | −57.4 | 107.56 | 71.93 | 11.92 | 57.64 |
| novel_miR_56 | TTTTGCCAGCTCCGCCCATTCC | 22 | 3p | 122 | 40.98 | −45.01 | 66.24 | 87.89 | 0 | 1.73 |
| novel_miR_57 | CAGCCAAGGATGACTTGCCGG | 21 | 5p | 175 | 35.43 | −57.4 | 0.97 | 1.66 | 0 | 0.92 |
| novel_miR_58 | TTTTTTAGTAGAAGGAGCAAAAT | 23 | 5p | 254 | 32.68 | −56.8 | 0.35 | 0 | 0 | 0 |
2.4. Expression Profiling of Differentially Expressed miRNAs in Response to V. dahliae Infection


2.5. Target Genes of miRNAs Identified by Degradome Analysis

| miRNA | Target Gene | Cleavage Site | Categoty | Norm Reads | Target Description | |
|---|---|---|---|---|---|---|
| Gb | Gh | |||||
| novel_miR_12 | Gorai.005G020900.1 | 1117 | 1 | 0 | 3 | Ubiquitin carboxyl-terminal hydrolase family protein |
| novel_miR_16 | Gorai.003G142500.1 | 1315 | 0 | 58 | 54 | Auxin response factor 10 isoform 1 |
| Gorai.013G267100.1 | 1416 | 0 | 60 | 31 | Auxin response factor 10 isoform 1 | |
| Gorai.010G046000.1 | 1638 | 0 | 16 | 19 | Auxin response factor 17 | |
| Gorai.006G166400.1 | 1773 | 0 | 36 | 12 | Auxin response factor 19 | |
| Gorai.011G238900.1 | 1502 | 0 | 316 | 557 | Auxin response factor 19 | |
| Gorai.012G004800.1 | 1679 | 0 | 110 | 160 | Auxin response factor 19 | |
| novel_miR_20 | Gorai.002G141500.1 | 618 | 4 | 1 | 0 | Purine permease 3 |
| Gorai.003G066400.1 | 1545 | 4 | 1 | 0 | Ferrochelatase 2 isoform 1 | |
| Gorai.008G199200.1 | 1792 | 4 | 1 | 0 | Uncharacterized protein TCM_014264 | |
| novel_miR_21 | Gorai.009G146400.1 | 1251 | 0 | 7 | 0 | Uncharacterized protein TCM_029129 |
| novel_miR_24 | Gorai.006G008600.5 | 496 | 0 | 1 | 0 | TT12-2 MATE transporter |
| novel_miR_26 | Gorai.007G279100.4 | 319 | 2 | 0 | 2 | Intracellular protein transport protein USO1 |
| Gorai.012G074700.1 | 79 | 2 | 0 | 2 | Serine hydroxymethyltransferase 6 isoform 3 | |
| novel_miR_28 | Gorai.003G163700.1 | 1250 | 0 | 13 | 0 | Leucine-rich repeat containing protein |
| novel_miR_29 | Gorai.008G010200.1 | 855 | 2 | 7 | 5 | Glutathione S-transferase 7 isoform 1 |
| Gorai.008G120500.1 | 1587 | 3 | 0 | 2 | Fringe-related protein, putative | |
| novel_miR_30 | Gorai.005G163400.1 | 1744 | 0 | 28 | 0 | Uncharacterized protein TCM_010813 |
| novel_miR_40 | Gorai.011G224600.3 | 1555 | 1 | 0 | 1 | Inositol 1,3,4-trisphosphate 5/6-kinase 4 |
| novel_miR_48 | Gorai.007G279100.4 | 319 | 2 | 0 | 2 | Intracellular protein transport protein USO1 |
| Gorai.012G074700.1 | 79 | 2 | 0 | 2 | Serine hydroxymethyltransferase 6 isoform 3 | |
| novel_miR_56 | Gorai.007G319800.1 | 613 | 0 | 10 | 1 | LRR and NB-ARC domains-containing disease resistance-like protein |
| novel_miR_57 | Gorai.004G172100.1 | 1219 | 0 | 19 | 26 | Nuclear transcription factor Y subunit A-1, putative isoform 1 |
| novel_miR_58 | Gorai.013G124500.1 | 2336 | 3 | 1 | 1 | Bacterial-induced lipoxygenase |
3. Discussion
4. Experimental Section
4.1. Plant Material and Total RNA Isolation
4.2. sRNA Library and Degradome Library Construction and Sequencing
4.3. Analysis of Sequencing Data
4.4. qRT-PCR
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Y.; Wang, W.; Chen, J.; Liu, J.; **a, M.; Shen, F. Identification of miRNAs and Their Targets in Cotton Inoculated with Verticillium dahliae by High-Throughput Sequencing and Degradome Analysis. Int. J. Mol. Sci. 2015, 16, 14749-14768. https://doi.org/10.3390/ijms160714749
Zhang Y, Wang W, Chen J, Liu J, **a M, Shen F. Identification of miRNAs and Their Targets in Cotton Inoculated with Verticillium dahliae by High-Throughput Sequencing and Degradome Analysis. International Journal of Molecular Sciences. 2015; 16(7):14749-14768. https://doi.org/10.3390/ijms160714749
Chicago/Turabian StyleZhang, Yujuan, Wei Wang, Jie Chen, Jubo Liu, Minxuan **a, and Fafu Shen. 2015. "Identification of miRNAs and Their Targets in Cotton Inoculated with Verticillium dahliae by High-Throughput Sequencing and Degradome Analysis" International Journal of Molecular Sciences 16, no. 7: 14749-14768. https://doi.org/10.3390/ijms160714749