Comparative Small RNA Analysis of Pollen Development in Autotetraploid and Diploid Rice
Abstract
:1. Introduction
2. Results
2.1. Overview of MicroRNAs (miRNAs) Sequencing Datasets in Develo** Pollens of Rice
2.2. Association between miRNAs Expression Profiles and Pollen Development Stages in Autotetraploid and Diploid Rice
2.3. Differentially Expressed miRNAs during Pollen Development in Autotetraploid and Diploid Rice
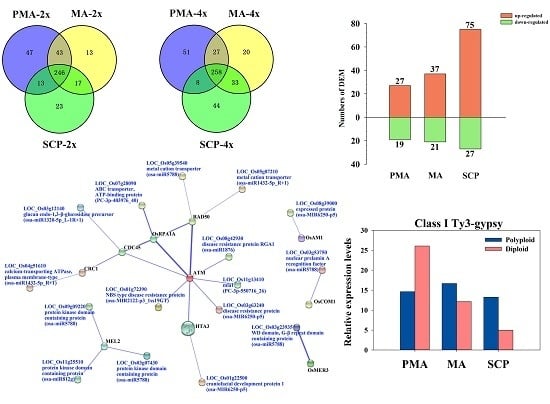
2.4. Target Prediction of Differentially Expressed miRNAs (DEM) and Functional Classification
2.5. Functional Classification of Meiosis-Related Predicted Targets of Differentially Expressed miRNAs
2.6. Functional Classification of Pre-Meiotic Interphase (PMA) and Single Microspore Stage (SCP) Stage-Specific Targets Regulated by Differentially Expressed miRNAs
2.7. Correlation between miRNAs and Their Target Genes
2.8. Differentially Expressed siRNAs in the Autopolyploid Pollen Associated with Transposable Elements
3. Discussion
3.1. Polyploidy Cause Changes in miRNA Expression Profiles during Pollen Development of Autotetraploid Rice
3.2. Specific miRNA Expression Patterns May Cause Meiosis Abnormalities in Autotetraploid Rice
3.3. Abundance of siRNAs Associated with Transposable Elements May Cause Meiosis Abnormalities in Autotetraploid Rice
4. Experimental Section
4.1. Rice Materials
4.2. Small RNA Library Construction, Sequencing and Data Processing
4.3. Analysis of Differentially Expressed miRNAs
4.4. The Prediction and Functional Analysis of Target Genes of miRNAs
4.5. Quantitative Real-Time PCR (qRT-PCR) Analysis
Supplementary Materials
Acknowledgments







| Material | Description/Stage-Specific Expression | Up-Regulated | Down-Regulated | Total |
|---|---|---|---|---|
| Taichung65-2x | DEM in MA compared to PMA | 20 | 46 | 66 |
| DEM in SCP compared to MA | 28 | 96 | 124 | |
| Taichung65-4x | DEM in MA compared to PMA | 41 | 75 | 116 |
| DEM in SCP compared to MA | 58 | 57 | 115 | |
| Taichung65-4x vs. Taichung65-2x | DEM in PMA | 27 | 19 | 46 |
| DEM in MA | 37 | 21 | 58 | |
| DEM in SCP | 75 | 27 | 102 | |
| DEM specifically in MA compared to PMA of Taichung65-2x | 15 | 34 | 49 | |
| DEM specifically in SCP compared to MA of Taichung65-2x | 19 | 55 | 74 | |
| DEM specifically in MA compared to PMA of Taichung65-4x | 36 | 63 | 99 | |
| DEM specifically in SCP compared to MA of Taichung65-4x | 49 | 16 | 65 |
| miRNA | Targets | Genes Annotation | PMA | MA | SCP | Specific in MA | Meiosis-Related Genes |
|---|---|---|---|---|---|---|---|
| osa-miR1320-5p_L-1R+1 | LOC_Os03g12140 | glucan endo-1,3-β-glucosidase precursor, putative, expressed | up | up | up | LOC_Os11g03430 (CDC45) | |
| osa-miR1432-5p_R+1 | LOC_Os04g51610 | calcium-transporting ATPase, plasma membrane-type, putative, expressed | up | up | up | LOC_Os04g40290 (CRC1) | |
| LOC_Os05g07210 | metal cation transporter, putative, expressed | up | up | up | LOC_Os02g29464 (RAD50) | ||
| osa-miR1876 | LOC_Os08g42930 | disease resistance protein RGA1, putative, expressed | ns | up | up | LOC_Os01g01689 (ATM) | |
| osa-MIR2122-p3_1ss19GT | LOC_Os01g72390 | NBS type disease resistance protein, putative, expressed | ns | up | ns | osa-MIR2122-p3_1ss19GT | LOC_Os01g01689 (ATM) |
| osa-miR5788 | LOC_Os03g07430 | protein kinase domain containing protein, expressed | ns | up | ns | osa-miR5788 | LOC_Os12g38460 (MEL2) |
| LOC_Os03g23935 | WD domain, G-β repeat domain containing protein, expressed | ns | up | ns | osa-miR5788 | LOC_Os02g40450 (OsMER3) | |
| LOC_Os03g53750 | nuclear prelamin A recognition factor, putative, expressed | ns | up | ns | osa-miR5788 | LOC_Os06g41050 (OsCOM1) | |
| LOC_Os05g39540 | metal cation transporter, putative, expressed | ns | up | ns | osa-miR5788 | LOC_Os02g29464 (RAD50) | |
| LOC_Os09g09220 | protein kinase domain containing protein, expressed | ns | up | ns | osa-miR5788 | LOC_Os12g38460 (MEL2) | |
| osa-MIR6250-p5 | LOC_Os01g22500 | craniofacial development protein 1, putative, expressed | ns | up | up | LOC_Os12g34510 (HTA3) | |
| LOC_Os03g63240 | disease resistance protein, putative, expressed | ns | up | up | LOC_Os01g01689 (ATM) | ||
| LOC_Os08g39000 | expressed protein | ns | up | up | LOC_Os03g44760 (OsAM1) | ||
| osa-miR812g | LOC_Os11g25510 | protein kinase domain containing protein, expressed | ns | up | up | LOC_Os12g38460 (MEL2) | |
| PC-3p-403976_40 | LOC_Os07g28090 | ABC transporter, ATP-binding protein, putative, expressed | ns | up | up | LOC_Os02g53680 (OsRPA1A) | |
| PC-3p-550716_26 | LOC_Os11g13410 | mla1, putative, expressed | ns | up | ns | PC-3p-550716_26 | LOC_Os01g01689 (ATM) |
| osa-miR160a-5p_R-1_1ss20CT | LOC_Os05g30820 | protein kinase, putative, expressed | ns | down | ns | osa-miR160a-5p_R-1_1ss20CT | LOC_Os12g38460 (MEL2) |
| osa-miR167a-3p_L+1R-1 | LOC_Os03g58260 | indole-3-glycerol phosphate lyase, chloroplast precursor, putative, expressed | ns | down | down | LOC_Os03g44760 (OsAM1) | |
| LOC_Os07g17220 | disease resistance protein, putative, expressed | ns | down | down | LOC_Os01g01689 (ATM) | ||
| osa-MIR5486-p3 | LOC_Os01g47256 | exosome complex exonuclease RRP40, putative, expressed | ns | down | down | LOC_Os01g72880 (MLH1) | |
| LOC_Os09g02700 | polyadenylate-binding protein, putative, expressed | ns | down | down | LOC_Os03g12414 (OsSDS) | ||
| LOC_Os11g39330 | NB-ARC domain containing protein, putative, expressed | ns | down | down | LOC_Os01g01689 (ATM) | ||
| osa-MIR5491-p5 | LOC_Os01g24060 | importin subunit α, putative, expressed | ns | down | ns | osa-MIR5491-p5 | LOC_Os03g12414 (OsSDS) |
| osa-MIR5518-p3 | LOC_Os04g48770 | ubiquitin fusion degradation protein, putative, expressed | ns | down | ns | osa-MIR5518-p3 | LOC_Os04g40290 (CRC1) |
| osa-MIR5806-p5 | LOC_Os08g19694 | NB-ARC domain containing protein, expressed | ns | down | ns | osa-MIR5806-p5 | LOC_Os01g01689 (ATM) |
| PC-3p-120829_194 | LOC_Os03g04520 | RNA recognition motif containing protein, putative, expressed | ns | down | ns | PC-3p-120829_194 | LOC_Os02g40450 (OsMER3) |
| MicroRNA_Family | miRNA | Sequence (5’ to 3’) | PMA | MA | SCP |
|---|---|---|---|---|---|
| miR2118 | osa-MIR2118k-p5 | TAAGCTCTATTGTCCCCTCTA | ns | ns | down |
| osa-miR2118e | TTCCCAATGCCTCCCATGCCTA | ns | ns | up | |
| osa-MIR2118l-p5 | TTAGGAAGAGGAAGAAATTGA | ns | down | ns | |
| osa-MIR2118m-p5 | GGAATGGGAACATGAAGGAAAG | ns | down | ns | |
| osa-MIR2118o-p5 | GGCATGGGGACATGAAGGAATG | ns | down | ns | |
| osa-miR2118a | TTCTCGATGCCTCCCATTCCTA | down | ns | ns | |
| osa-miR2118c | TTCCCGATGCCTCCTATTCCTA | down | ns | ns | |
| osa-MIR2118a-p5 | GGACTGGGAACATATGAGAAAG | down | ns | ns | |
| miR2275 | osa-miR2275d | CTTGTTTTTCTCCAATATCTCA | down | ns | up |
| osa-MIR2275c-p3 | ATTGTTTTTCTCCAATATCTCA | down | ns | up | |
| mes-miR2275 | TTTGGTTTCCTCCAATATCTTA | down | ns | ns | |
| zma-miR2275b-3p_1ss22AG | TTCAGTTTCCTCTAATATCTCG | down | ns | ns |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, X.; Shahid, M.Q.; Wu, J.; Wang, L.; Liu, X.; Lu, Y. Comparative Small RNA Analysis of Pollen Development in Autotetraploid and Diploid Rice. Int. J. Mol. Sci. 2016, 17, 499. https://doi.org/10.3390/ijms17040499
Li X, Shahid MQ, Wu J, Wang L, Liu X, Lu Y. Comparative Small RNA Analysis of Pollen Development in Autotetraploid and Diploid Rice. International Journal of Molecular Sciences. 2016; 17(4):499. https://doi.org/10.3390/ijms17040499
Chicago/Turabian StyleLi, **ang, Muhammad Qasim Shahid, **wen Wu, Lan Wang, **angdong Liu, and Yonggen Lu. 2016. "Comparative Small RNA Analysis of Pollen Development in Autotetraploid and Diploid Rice" International Journal of Molecular Sciences 17, no. 4: 499. https://doi.org/10.3390/ijms17040499