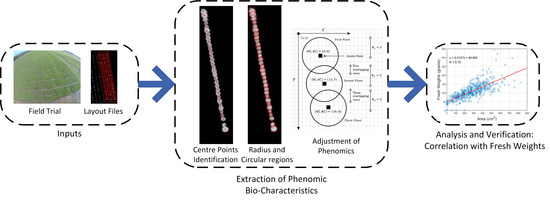
A New Method for Extracting Individual Plant Bio-Characteristics from High-Resolution Digital Images
Abstract
:1. Introduction
2. Problem Statements
- to identify bounding boxes with no plants;
- to calculate accurate individual plant areas, despite overlap** adjacent plants;
- to calculate accurate individual plant NDVI values, despite overlap** adjacent plants;
3. Methods
3.1. Background Correction
3.2. Center Point Calculation
3.3. Extraction of Plant Areas
3.4. Extraction of NDVI Values
3.4.1. Finding the Overlap** Pixel Rows
3.4.2. Adjusting NDVI Values at Overlap** Pixel Rows
- the maximum and minimum NDVI values of the plant are first calculated, labelled as and ; respectively;
- the whole center row is updated and will be used as a reference for the adjustment of plant pixels at overlap** rows. The step size, which is the difference of NDVI values between two adjacent pixels, is calculated as:
- Let us take a symmetric reference vector, , such that The NDVI values are adjusted as following:
3.5. Testing of the Algorithm
4. Results and Discussions
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Tester, M.; Langridge, P. Breeding Technologies to Increase Crop Production in a Changing World. Science 2010, 327, 818–822. [Google Scholar] [CrossRef]
- Long, S.P.; Marshall-Colon, A.; Zhu, X.-G. Meeting the Global Food Demand of the Future by Engineering Crop Photosynthesis and Yield Potential. Cell 2015, 161, 56–66. [Google Scholar] [CrossRef] [Green Version]
- Brown, T.B.; Cheng, R.; Sirault, X.R.R.; Rungrat, T.; Murray, K.D.; Trtilek, M.; Furbank, R.T.; Badger, M.; Pogson, B.J.; O Borevitz, J. TraitCapture: Genomic and environment modelling of plant phenomic data. Curr. Opin. Plant Biol. 2014, 18, 73–79. [Google Scholar] [CrossRef] [Green Version]
- Kumar, J.; Pratap, A.; Kumar, S. Plant Phenomics: An Overview. In Phenomics in Crop Plants: Trends, Options and Limitations; Springer: New Delhi, India, 2015; pp. 1–10. [Google Scholar]
- Sticklen, M.B. Feedstock Crop Genetic Engineering for Alcohol Fuels. Crop. Sci. 2007, 47, 2238–2248. [Google Scholar] [CrossRef] [Green Version]
- Van der Kooi, C.J.; Reich, M.; Löw, M.; De Kok, L.J.; Tausz, M. Growth and yield stimulation under elevated CO2 and drought: A meta-analysis on crops. Environ. Exp. Bot. 2017, 122, 150–157. [Google Scholar] [CrossRef]
- O’Malley, R.C.; Ecker, J.R. Linking genotype to phenotype using the Arabidopsis unimutant collection. Plant J. 2010, 61, 928–940. [Google Scholar] [CrossRef]
- Weigel, D.; Richard, M. The 1001 Genomes Project for Arabidopsis thaliana. Genome Biol. 2009, 10, 1–5. [Google Scholar] [CrossRef] [Green Version]
- Cannon, S.B.; May, G.D.; Jackson, S.A. Three Sequenced Legume Genomes and Many Crop Species: Rich Opportunities for Translational Genomics. Plant Physiol. 2009, 151, 970–977. [Google Scholar] [CrossRef] [Green Version]
- International Brachypodium Initiative. Genome sequencing and analysis of the model grass Brachypodium distachyon. Nature 2011, 463, 763–770. [Google Scholar]
- Atwell, S.; Huang, Y.S.; Vilhjálmsson, B.J.; Willems, G.; Horton, M.W.; Li, Y.; Meng, D.; Platt, A.; Tarone, A.M.; Hu, T.T.; et al. Genome-wide association study of 107 phenotypes in Arabidopsis thaliana inbred lines. Nat. Cell Biol. 2010, 465, 627–631. [Google Scholar] [CrossRef]
- Wang, M.; Jiang, N.; Jia, T.; Leach, L.; Cockram, J.; Comadran, J.; Shaw, P.; Waugh, R.; Luo, Z. Genome-wide association map** of agronomic and morphologic traits in highly structured populations of barley cultivars. Theor. Appl. Genet. 2012, 124, 233–246. [Google Scholar] [CrossRef] [PubMed]
- Lucocq, J.M. Efficient quantitative morphological phenoty** of genetically altered organisms using stereology. Transgenic Res. 2007, 16, 133–145. [Google Scholar] [CrossRef] [PubMed]
- Chung, K.; Crane, M.M.; Lu, H. Automated on-chip rapid microscopy, phenoty** and sorting of C. elegans. Nat. Methods 2008, 5, 637–643. [Google Scholar] [CrossRef] [PubMed]
- Sozzani, R.; Benfey, P.N. High-throughput phenoty** of multicellular organisms: Finding the link between genotype and phenotype. Genome Biol. 2011, 12, 219. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Souza, N. High-throughput phenoty**. Nat. Methods 2010, 7, 1. [Google Scholar] [CrossRef]
- P-Sanz, F.; Navarro, P.J.; E-Cortines, M. Plant phenomics: An overview of image acquisition technologies and image data analysis algorithms. Gigascience 2017, 6, 1–18. [Google Scholar]
- Badrinarayanan, V.; Kendall, A.; Cipolla, R. SegNet: A Deep Convolutional Encoder-Decoder Architecture for Image Segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 39, 2481–2495. [Google Scholar] [CrossRef] [PubMed]
- Otsu, N. A threshold selection method from gray-level histograms. IEEE Trans. Syst. Man, Cybern. 1979, 9, 62–66. [Google Scholar] [CrossRef] [Green Version]
- Vincent, L.; Soille, P. Watersheds in digital spaces: An efficient algorithm based on immersion simulations. IEEE Trans. Pattern Anal. Mach. Intell. 1991, 13, 583–598. [Google Scholar] [CrossRef] [Green Version]
- Banerjee, B.P.; Spangenberg, G.; Kant, S. Fusion of Spectral and Structural Information from Aerial Images for Improved Biomass Estimation. Remote Sens. 2020, 12, 3164. [Google Scholar] [CrossRef]
- Gebremedhin, A.; Badenhorst, P.; Wang, J.; Shi, F.; Breen, E.; Giri, K.; Spangenberg, G.C.; Smith, K. Development and Validation of a Phenoty** Computational Workflow to Predict the Biomass Yield of a Large Perennial Ryegrass Breeding Field Trial. Front. Plant Sci. 2020, 11. [Google Scholar] [CrossRef]
- Silva, J.C.E.; Kerr, R.J. Accounting for competition in genetic analysis, with particular emphasis on forest genetic trials. Tree Genet. Genomes 2012, 9, 1–17. [Google Scholar] [CrossRef]
- Tucker, C.J. Red and photographic infrared linear combinations for monitoring vegetation. Remote Sens. Environ. 1979, 8, 127–150. [Google Scholar] [CrossRef] [Green Version]
- Neumann, K.; Klukas, C.; Friedel, S.; Rischbeck, P.; Chen, D.; Entzian, A.; Stein, N.; Graner, A.; Kilian, B. Dissecting spatiotemporal biomass accumulation in barley under different water regimes using high-throughput image analysis. Plant Cell Environ. 2015, 38, 1980–1996. [Google Scholar] [CrossRef]
- Tackenberg, O. A New Method for Non-destructive Measurement of Biomass, Growth Rates, Vertical Biomass Distribution and Dry Matter Content Based on Digital Image Analysis. Ann. Bot. 2006, 99, 777–783. [Google Scholar] [CrossRef]
- Chen, D.; Neumann, K.; Friedel, S.; Kilian, B.; Chen, M.; Altmann, T.; Klukas, C. Dissecting the Phenotypic Components of Crop Plant Growth and Drought Responses Based on High-Throughput Image Analysis. Plant Cell 2014, 26, 4636–4655. [Google Scholar] [CrossRef] [Green Version]
- Hartmann, A.; Czauderna, T.; Hoffmann, R.; Stein, N.; Schreiber, F. HTPheno: An image analysis pipeline for high-throughput plant phenoty**. BMC Bioinform. 2011, 12, 148. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Subramanian, R.; Spalding, E.P.; Ferrier, N.J. A high throughput robot system for machine vision based plant phenotype studies. Mach. Vis. Appl. 2012, 24, 619–636. [Google Scholar] [CrossRef] [Green Version]
- Miller, N.D.; Parks, B.M.; Spalding, E.P. Computer-vision analysis of seedling responses to light and gravity. Plant J. 2007, 52, 374–381. [Google Scholar] [CrossRef]
- Clark, R.T.; MacCurdy, R.B.; Jung, J.K.; Shaff, J.E.; McCouch, S.R.; Aneshansley, D.J.; Kochian, L.V. Three-Dimensional Root Phenoty** with a Novel Imaging and Software Platform. Plant Physiol. 2011, 156, 455–465. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Atherton, T.; Kerbyson, D. Size invariant circle detection. Image Vis. Comput. 1999, 17, 795–803. [Google Scholar] [CrossRef]
- Yuen, H.K.; Princen, J.; Dlingworth, J.; Kittler, J. A Comparative Study of Hough Transform Methods for Circle Finding. Image Vis. Comput. 1989, 8, 71–77. [Google Scholar] [CrossRef] [Green Version]
- Gebremedhin, A.; Badenhorst, P.; Wang, J.; Giri, K.; Spangenberg, G.; Smith, K. Development and Validation of a Model to Combine NDVI and Plant Height for High-Throughput Phenoty** of Herbage Yield in a Perennial Ryegrass Breeding Program. Remote Sens. 2019, 11, 2494. [Google Scholar] [CrossRef] [Green Version]
- Rodgers, J.L.; Nicewander, W.A. Thirteen Ways to Look at the Correlation Coefficient. Am. Stat. 1988, 42, 59. [Google Scholar] [CrossRef]
- Daetwyler, H.D.; Villanueva, B.; Woolliams, J.A. Accuracy of Predicting the Genetic Risk of Disease Using a Genome-Wide Approach. PLoS ONE 2008, 3, e3395. [Google Scholar] [CrossRef] [PubMed] [Green Version]













| Image Time Point | Correlation | |
|---|---|---|
| Area of Rectangular Bounding Boxes | Area of Circular Plant Regions | |
| 9 May 2017 | 0.74 | 0.75 |
| 5 July 2017 | 0.30 | 0.74 |
| 11 September 2017 | 0.28 | 0.63 |
| 20 November 2017 | 0.30 | 0.66 |
| Image Time Point | Correlation | ||
|---|---|---|---|
| Unadjusted NDVI from Rectangular Boxes | Unadjusted NDVI from Circular Plant Regions | Adjusted NDVI from Circular Plant Regions | |
| 9 May 2017 | 0.56 | 0.56 | 0.57 |
| 5 July 2017 | 0.55 | 0.58 | 0.59 |
| 11 September 2017 | 0.52 | 0.54 | 0.55 |
| 20 November 2017 | 0.51 | 0.53 | 0.56 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rabab, S.; Breen, E.; Gebremedhin, A.; Shi, F.; Badenhorst, P.; Chen, Y.-P.P.; Daetwyler, H.D. A New Method for Extracting Individual Plant Bio-Characteristics from High-Resolution Digital Images. Remote Sens. 2021, 13, 1212. https://doi.org/10.3390/rs13061212
Rabab S, Breen E, Gebremedhin A, Shi F, Badenhorst P, Chen Y-PP, Daetwyler HD. A New Method for Extracting Individual Plant Bio-Characteristics from High-Resolution Digital Images. Remote Sensing. 2021; 13(6):1212. https://doi.org/10.3390/rs13061212
Chicago/Turabian StyleRabab, Saba, Edmond Breen, Alem Gebremedhin, Fan Shi, Pieter Badenhorst, Yi-** Phoebe Chen, and Hans D. Daetwyler. 2021. "A New Method for Extracting Individual Plant Bio-Characteristics from High-Resolution Digital Images" Remote Sensing 13, no. 6: 1212. https://doi.org/10.3390/rs13061212