1. Introduction
Anal, vulvar, and penile cancer are tumors of external genitalia considered as rare due to their low incidence among the global population, showing an incidence of 0.65, 1.2 and 0.92 cases per 100,000 habitants in 2020, respectively [
1,
2]. In Chile, these types of cancer are also rare, with between 100 and 140 new cases diagnosed per year. During 2020, the cases of anal, vulvar and penile cancer were 134, 128 and 93, respectively, with a prevalence at five years of 1.8, 3.5, and 2.6 per 100,000 Chileans [
1].
All of these tumors have in common their development from a premalignant lesion related to the Human Papilloma Virus (HPV) infection, and in some cases arising through an HPV independent pathway [
3,
4].
HPV oncoproteins can induce genetic alterations affecting cell proliferation, the cell cycle, and genomic stability [
5,
6,
7]. In addition, HPV has been proven to induce alterations in the immune microenvironment of infected tissue such as the suppression of immune recognition, promoting acute inflammation and the upregulation of immune checkpoint proteins [
8,
9,
10].
The most common treatment for these rare cancers is chemo-radiotherapy and surgery; however, several other strategies are now in clinical trials. As with many other types of cancer, immunotherapy has also been evaluated as a treatment option [
10,
11,
12,
13], therefore a comprehensive characterization of tissue biomarkers within the tumor microenvironment of rare cancers is essential.
Tumor-infiltrating lymphocytes (TILs) have shown a crucial role in the anti-tumor immune response as have macrophages, and penile, vulvar, and anal cancer are no exception, with several groups describing the presence of these markers and their role in outcome and survival [
9,
14,
15,
16,
17,
18,
19,
20,
21]. On the other hand, tumor cells can evade immune anti-tumor response using several mechanisms, such as the expression of negative immune checkpoint molecules and the modulation of the tumor microenvironment [
22,
23,
24,
25]. The expression of PD-L1 has been demonstrated in penile [
26,
27], vulvar [
28,
29] and anal [
30,
31] carcinomas, with different percentages of positive cells, and these previous reports about PD-L1 expression, including TILs presence, in rare cancer tissues had been evaluated using a classic immunohistochemistry (IHC) [
26,
27,
28,
29,
30,
31]. Even though the IHC is considered as a hallmark for cancer diagnosis, this tool has several limitations, including inter-observer variability, and the fact that it only allows the analysis of one to two biomarkers at a time [
32]. Hence, in order to analyze several markers, several histological sections are required, which is challenging when only a small sample of tissue is available [
33], or when the cancer type of interest is classified as rare and the tissue sample is limited by the frequency of appearance.
Several tools have been developed for multiplexing targets during histological studies [
34]; however, the least expensive is based on multiplexed immunofluorescence (mIF), which advances those challenges by interrogating panels of multiple biomarkers with 5–6 being the most common and up to 9 depending on the instruments and methodology. Due to this advantage, the mIF analysis is a useful tool to characterize the tumor microenvironment, particularly in tumor types with low incidences such as penile, vulvar and anal cancer, and a limited amount of tissue for further studies.
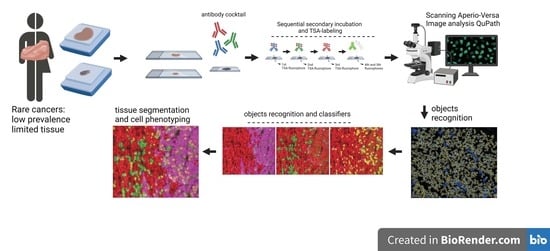
In this study we describe the technical approach during the implementation of two panels of validated antibodies to evaluate the proliferation rate of tumor cells based on Ki67 signal, tumor infiltrating immune cells by CD8 signal, and by CD68 detection for macrophages, and the tissue localization of the immune checkpoint PD-L1, applying tissue segmentation and cell phenoty** to interrogate the cytokeratin positive and negative compartments in retrospective cases of rare cancer in Chilean patients.
2. Materials and Methods
2.1. Patients and Case Selection
This is a single-institution retrospective study at Instituto Oncológico Fundación Arturo López Pérez (Santiago, Chile), reviewed and approved by the institutional ethical committee (Prot. 007-RES-ANC-MUL). Deceased patients with histologically proven invasive squamous cell carcinoma of penis, vulva and anus diagnosed in the Institution during 2010–2014 who died because of these diseases between 2014 and 2017 were considered eligible. The respective historical FFPE block was collected from the internal biorepository after the Institutional Ethical Committee approval of the waiver of a signed consent letter. Twenty patient records were retrospectively found, however four cases of penile cancer, one case of vulvar cancer and two of anal cancer had complete clinical records and a suitable tissue sample (
Table 1). Clinical and demographic data were retrieved from patient medical records and managed according to privacy regulations.
2.2. Sample Processing and Multiplexed Immunofluorescence
For each tumor type, all available tissue cores were requested and examined by a pathologist to confirm the histology and the integrity of the material. Based on hematoxylin-eosin visual inspection, all cores with an area of less than 5% of the tumor were excluded. Immunostaining was carried out as published [
35] and each primary cocktail (or panel) was developed using validated antibodies [
35,
36]. In panel 1: PD-L1 (clone E1L3N), Cytokeratin (clone AE1/AE3), CD68 (clone PG-M1) and CD8a (clone C8/144B). In panel 2: PD-1 (clone D4W2J), Cytokeratin (clone AE1/AE3) and Ki67 (clone MIB-1). Briefly, after antigen retrieval, each whole tissue slide was incubated overnight at 4 °C with the selected panel separately. Isotype-specific HRP-conjugated antibodies and tyramide-based amplification systems (Perkin Elmer, Waltham, MA, USA) were used for signal detection. After co-staining with DAPI, each slide was cover slipped in Prolong Gold mounting medium (Invitrogen, Carlsbad, CA, USA). An index TMA was stained in parallel as quality control. This array included human tissue as naturally positive and negative controls, such as skeletal muscle (negative/noise threshold), tonsil (positive for immune cells), and placenta (positive for PD-L1 immune checkpoint). The dynamic range of positivity for each biomarker was estimated from the ratio between the mean fluorescent signal from a positive control tissue and a negative control tissue. Pearson’s coefficients were calculated between independent staining runs or for single markers to verify concordances between experiments.
2.3. Cases and FFPE Cores Considerations
For some analysis considering average values in penile cancer, only cases with more than one FFPE core were considered, however for single case representation all cases were included. For the single case of vulvar cancer, the three FFPE cores available were processed and analyzed in some figures as average values or individual values. For anal cancer, each case and its respective FFPE core was analyzed separately.
2.4. Slide Scanning and Image Generation
All slides were scanned at 20× of magnification using an Aperio VERSA 200 microscope (Leica Biosystems, Vista, CA, USA). Exposure times were determined for each target using an index TMA including normal human tissue as positive and negative controls, such as skeletal muscle, tonsil and placenta. The optimal exposure time considered the optimal signal to noise ratio measured for each target and the supervised observation of the operator for the visual positive signal and the expected staining pattern. Later, for each antibody panel, an unsupervised template was designed for the automatic and simultaneous scanning of the fluorescent signal and applied to the tumor whole tissue section. The templates considered the following steps: an initial low-resolution scanning at 1.25× for automatic tissue detection, a second scanning at 5× for fluorescent signal detection (using DAPI); and a final scanning at 20× including sequentially FITC, Cy3, Cy5 and Cy7 filters. Images were saved in scn extension and size varied from 26,976 × 32,249 to 60,440 × 32,249 pixels. After acquisition, all images were revised and exported for further analysis.
2.5. Analysis for Single Objects
Whole-slide scans from the fluorescent-labeled sections were analyzed using QuPath v.0.3.2 software (University of Edinburgh. Edinburgh, Scotland, UK) [
37]. DAPI channel was used for cell segmentation using the following settings: Pixel size: 0.5 μm; Background radius: 8.0 μm; Median filter radius: 0.0 μm; Minimum area: 10.0 μm
2; Maximum area: 200 μm
2; Threshold: 5.0.
After cell detection, cells were classified using the Object classifier module to determine the number of positive cells for each marker after applying the defined visual cutoff (fluorescent intensity ratio). At least 400 cells from different cores were used to train each object classifier. After every object classifier was trained, they were run for each image using the script editor for batch analysis. Data was exported and analyzed using Microsoft Office tools. The number of total positive cells for each biomarker was determined by the addition of the positive cells for individual biomarkers and the positive ones for more than one biomarker. Percentages were calculated based on the total cell detections.
2.6. Analysis for Segmentation and Masking
The pixel classifier from QuPath was trained under manual supervision to detect four different regions: Ignore, corresponding to the sections of the image that had no tissue (glass); Negative, corresponding to regions with auto-fluorescence, mostly red blood cells; Tumor region, defined by Cytokeratin and DAPI positive signal, and Stroma region, corresponding to areas without signal for Cytokeratin, but including DAPI positive objects. For the pixel classifier, 28 fields of view from different tissue images were generated and used for the image training, and the trained classifier was applied in batch or as a single run to each digitized image. The automatic tumor masking was verified visually based on nuclei morphology. Once Pixel classifiers were verified, the tumor and stromal areas were set as annotations and cells within each annotation (tumor or stroma) were classified using the Object Classifier plugin. Data was exported and analyzed in Microsoft Excel. Percentages were calculated based on the total cell detections.
4. Discussion
The Society for Immunotherapy of Cancer (SITC) has provided the fundamentals of best practices for multiplex immunohistochemistry (mIHC) and immunofluorescence (mIF) staining and validation, and suggested that mIHC/mIF technologies are becoming standard tools for biomarker studies and are likely to enter routine clinical practice in the near future [
38]. For that reason, a careful assay optimization and validation is required to ensure that outputs are robust and comparable across laboratories as well as potentially across mIHC/mIF platforms.
This study represents the beginning of the use of multiplexing and its validation process developed by a Chilean cancer center based on the recommendations of the SITC, and included an objective and reliable approach for the characterization of a tumor microenvironment in FPEE samples from invasive squamous cell carcinomas of the penis, vulva and anus. These types of tumors are classified as rare for their low incidence in the population compared to other types of malignancies [
1,
39,
40]. Since the number of samples is always limited for this type of cancer, we have developed a multiplex immunofluorescence analysis that allowed us to obtain more information from a single histological section.
After reviewing three different case series of two anal cancers (single tissue core each), of one vulvar cancer (three tissue cores), and of four penile cancers (two cores and one single core), using a single biomarker for cell phenoty** may reduce the possibility for the researcher and the pathologist to detect the natural variability between cores and cases, and the combination of biomarkers may highlight the differences between patients and even between tissue cores from a same patient.
As far as we know, this is one of the first studies to analyze FFPE samples from rare cancers in Chilean patients by multiplexed immunofluorescence (mIF). Previously, studies based on image analysis from FFPE samples from these types of cancer were performed using classic IHC, which has a limitation regarding the number of biomarkers to analyze and a high inter-observer variability [
32,
33].
The mIF facilitates the study of these rare cancers because it enables us to analyze the co-expression of different biomarkers for a better understanding of the tumor microenvironment, identifying different cell phenotypes combining biomarkers and maintaining the spatial distribution for each detected element [
41]. Using an open-source software such as QuPath allows anyone, including research laboratories with limited resources, to collect a large amount of data, from single counts, location and distribution and regional abundance using positive pixel training to identify the target of interest [
42,
43,
44,
45]. Based on the software’s plugins, we analyzed the number of positive cells for each biomarker in the whole tissue section and ran the analysis in batch, which reduced time and variability inter-reader [
45,
46]. After managing the single cell identifier, larger areas of tissue can be analyzed by specific signal masking, such as tumor nest and stroma. The QuPath plugin, through a supervised training, identifies pixels with a pan-cytokeratin antibody signal, segregating them into pixels with and without information to define an algorithm for the automatic tissue masking and the execution of a secondary analysis, such as counting single elements within the selected area as a regular machine learning operation [
47,
48].
The selection of biomarkers used in this work was based on their clinical relevance and role in tumor microenvironment adaptation. Tumor-infiltrating CD8 lymphocytes are considered as the major inflammatory component in almost all known solid tumors and are responsible for killing tumor cells by their cytotoxic activity [
49,
50] In addition, these TILs may become exhausted because they are exposed to a sustained immune suppressive environment, such as tumor cells and myeloid cells expressing PD-L1. As reported by our collaborators and other researchers, using immune checkpoint blockers may reinvigorate some cytotoxic CD8 T cells and restore their effector role against several tumor types [
51,
52]. In addition, radiotherapy is frequently used to treat these type of cancers, and recently an abscopal effect after irradiation regimens was described in preclinical models where CD8 T cells were shown to be crucial in improving the response to immunomodulatory monoclonal antibodies, such as anti-PD1 or anti-CD137 [
52].
Different mechanisms of immune evasion have been studied in penile [
11,
21,
53,
54] vulvar [
14,
18,
20,
55] and anal cancers [
17,
19], and included immune checkpoint proteins and exhausted phenotypes for infiltrating-lymphocytes, suggesting that patients with these type of tumors can benefit from anti PD-L1 therapy [
9,
27,
56,
57,
58,
59]. Currently, there are several clinical trials ongoing for the use of immune therapy targeting PD-L1 (or its receptor PD-1) as treatment for penile, vulvar or anal cancer [
60,
61,
62,
63,
64,
65,
66,
67,
68,
69]. Therefore, multiplexed immunofluorescence and an objective analysis of the tumor microenvironment may contribute significantly to improve the understanding of different therapeutic targets when the access to biological material is reduced or limited.
Flow cytometry has been the most extended tool to characterize specific cell phenotypes based on the multiple detection of surface and intracellular biomarkers, in particular for non-tumor cells within the tumor microenvironment, such as infiltrating lymphocytes [
70] and tumor-associated macrophages (TAM) [
71]. We did not include specific phenotypes, such as M1 or M2 macrophages, because of the lack of validated antibodies for histochemistry or the complexity of combining some of them, and CD68 alone may be considered as an indicator of TAM-like cells. Other limitations of this study include the small number of analyzed cases which may affect the ability of the trained algorithms to recognize a variable related to a specific-tissue type, in particular in the only one case of vulvar cancer. In addition, we were aware of the absence of statistical power, which limited the analysis but it does not invalidate the histological findings and developed tools. Finally, we determined the localization of single objects but we did not carry out algorithms to measure distance between objects, which has also been described as being determinant in tumor microenvironment characterization [
72].