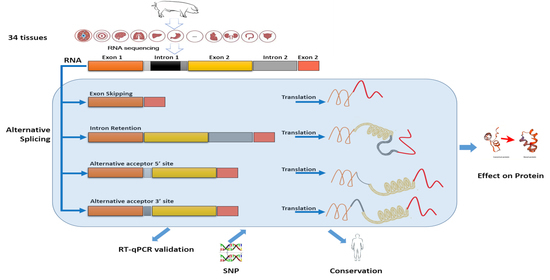
Profiling Novel Alternative Splicing within Multiple Tissues Provides Useful Insights into Porcine Genome Annotation
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sample Collection and Sequencing
2.2. Separation of RNA from Tissues
2.3. Library Construction and RNA Sequencing
2.4. Read Alignment to the Reference Sus Scrofa 11.1 Genome
2.5. Alternative Splicing Events in Pig Transcriptome
2.6. Comparison of Novel and Known Protein Domains
2.7. SNP Compared with Alternative Splicing Variations
2.8. Known and Novel AS Validation Using Quantitative Reverse Transcription PCR
2.9. Protein Extraction and Western Blotting
3. Results
3.1. Identification of Novel Transcripts in 34 Different Pig Tissues
3.2. Classification of Alternative Splicing Types
3.3. Tissue Specificity of Alternative Splicing in Different Tissues
3.4. Potential Effect of Novel Alternative Splicing on the Pig Proteome
3.5. Conserved Alternative Splicing between Pig and Human
3.6. Effect of Single Nucleotide Polymorphism on Pig Alternative Splicing
3.7. Validation of Known and Novel Transcripts Using RT-qPCR and Western Blotting
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Pan, Q.; Shai, O.; Lee, L.J.; Frey, B.J.; Blencowe, B.J. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat. Genet. 2008, 40, 1413–1415. [Google Scholar] [CrossRef] [PubMed]
- Chacko, E.; Ranganathan, S. Genome-wide analysis of alternative splicing in cow: Implications in bovine as a model for human diseases. BMC Genom. 2009, 10, S11. [Google Scholar] [CrossRef] [Green Version]
- Liu, S.; Zhou, X.; Hao, L.; Piao, X.; Hou, N.; Chen, Q. Genome-Wide Transcriptome Analysis Reveals Extensive Alternative Splicing Events in the Protoscoleces of Echinococcus granulosus and Echinococcus multilocularis. Front. Microbiol. 2017, 8, 929. [Google Scholar] [CrossRef] [PubMed]
- Sammeth, M.; Foissac, S.; Guigo, R. A General Definition and Nomenclature for Alternative Splicing Events. PLoS Comput. Biol. 2008, 4, e1000147. [Google Scholar] [CrossRef] [Green Version]
- Koenigsberger, C.; Chicca, J.J.; Amoureux, M.C.; Edelman, G.M.; Jones, F.S. Differential regulation by multiple promoters of the gene encoding the neuron-restrictive silencer factor. Proc. Natl. Acad. Sci. USA 2000, 97, 2291–2296. [Google Scholar] [CrossRef] [Green Version]
- Di Giammartino, D.C.; Nishida, K.; Manley, J.L. Mechanisms and Consequences of Alternative Polyadenylation. Mol. Cell 2011, 43, 853–866. [Google Scholar] [CrossRef] [Green Version]
- Emrich, S.J.; Barbazuk, W.B.; Li, L.; Schnable, P.S. Gene discovery and annotation using LCM-454 transcriptome sequencing. Genome Res. 2007, 17, 69–73. [Google Scholar] [CrossRef] [Green Version]
- Ruhl, C.; Stauffer, E.; Kahles, A.; Wagner, G.; Drechsel, G.; Ratsch, G.; Wachter, A. Polypyrimidine Tract Binding Protein Homologs from Arabidopsis Are Key Regulators of Alternative Splicing with Implications in Fundamental Developmental Processes. Plant. Cell 2012, 24, 4360–4375. [Google Scholar] [CrossRef] [Green Version]
- Staiger, D.; Brown, J.W.S. Alternative Splicing at the Intersection of Biological Timing, Development, and Stress Responses. Plant. Cell 2013, 25, 3640–3656. [Google Scholar] [CrossRef] [Green Version]
- Gutierrez-Arcelus, M.; Ongen, H.; Lappalainen, T.; Montgomery, S.B.; Buil, A.; Yurovsky, A.; Bryois, J.; Padioleau, I.; Romano, L.; Planchon, A.; et al. Tissue-Specific Effects of Genetic and Epigenetic Variation on Gene Regulation and Splicing. PLoS Genet. 2015, 11, e1004958. [Google Scholar] [CrossRef] [Green Version]
- Baralle, F.E.; Giudice, J. Alternative splicing as a regulator of development and tissue identity. Nat. Rev. Mol. Cell Bio. 2017, 18, 437–451. [Google Scholar] [CrossRef]
- Scotti, M.M.; Swanson, M.S. RNA mis-splicing in disease. Nat. Rev. Genet. 2016, 17, 19–32. [Google Scholar] [CrossRef]
- Duque, P. A role for SR proteins in plant stress responses. Plant Signal. Behav. 2011, 6, 49–54. [Google Scholar] [CrossRef] [Green Version]
- Sun, X.Y.; Yang, Q.Y.; Deng, Z.P.; Ye, X.F. Digital Inventory of Arabidopsis Transcripts Revealed by 61 RNA Sequencing Samples. Plant Physiol. 2014, 166, 869–878. [Google Scholar] [CrossRef] [Green Version]
- Cheng, J.H.; Zhou, T.; Liu, C.D.; Shapiro, J.P.; Brauer, M.J.; Kiefer, M.C.; Barr, P.J.; Mountz, J.D. Protection from Fas-Mediated Apoptosis by a Soluble Form of the Fas Molecule. Science 1994, 263, 1759–1762. [Google Scholar] [CrossRef]
- Jiang, F.K.; Guo, M.; Yang, F.; Duncan, K.; Jackson, D.; Rafalski, A.; Wang, S.C.; Li, B.L. Mutations in an AP2 Transcription Factor-Like Gene Affect Internode Length and Leaf Shape in Maize. PLoS ONE 2012, 7, e37040. [Google Scholar] [CrossRef] [Green Version]
- Li, X.R.; Zhu, C.S.; Yeh, C.T.; Wu, W.; Takacs, E.M.; Petsch, K.A.; Tian, F.; Bai, G.H.; Buckler, E.S.; Muehlbauer, G.J.; et al. Genic and nongenic contributions to natural variation of quantitative traits in maize. Genome Res. 2012, 22, 2436–2444. [Google Scholar] [CrossRef] [Green Version]
- ** events differentiate among splicing patterns in sixteen human tissues. F1000Research 2013, 2, 188. [Google Scholar] [CrossRef]
- Kroll, J.E.; Kim, J.; Ohno-Machado, L.; de Souza, S.J. Splicing Express: A software suite for alternative splicing analysis using next-generation sequencing data. PeerJ 2015, 3, e1419. [Google Scholar] [CrossRef] [Green Version]
- HMMER3. Available online: https://www.ebi.ac.uk/Tools/hmmer/ (accessed on 9 August 2018).
- Swiss-Model. Available online: https://swissmodel.expasy.org/ (accessed on 2 July 2020).
- Wang, L.; Nie, J.J.; Kocher, J.P. PVAAS: Identify variants associated with aberrant splicing from RNA-seq. Bioinformatics 2015, 31, 1668–1670. [Google Scholar] [CrossRef] [Green Version]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Buchfink, B.; **e, C.; Huson, D.H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 2015, 12, 59–60. [Google Scholar] [CrossRef]
- Muvarak, N.; Kelley, S.; Robert, C.; Baer, M.R.; Perrotti, D.; Gambacorti-Passerini, C.; Civin, C.; Scheibner, K.; Rassool, F.V. c-MYC Generates Repair Errors via Increased Transcription of Alternative-NHEJ Factors, LIG3 and PARP1, in Tyrosine Kinase–Activated Leukemias. Mol. Cancer Res. 2015, 13, 699–712. [Google Scholar] [CrossRef] [Green Version]
- Wang, H.; Chen, Y.; Li, X.; Chen, G.; Zhong, L.; Chen, G.; Liao, Y.; Liao, W.; Bin, J. Genome-wide analysis of alternative splicing during human heart development. Sci. Rep. 2016, 6, 35520. [Google Scholar] [CrossRef] [Green Version]
- Sheth, N.; Roca, X.; Hastings, M.L.; Roeder, T.; Krainer, A.R.; Sachidanandam, R. Comprehensive splice-site analysis using comparative genomics. Nucleic Acids Res. 2006, 34, 3955–3967. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, J.; Jiang, H.; **e, T.; Zheng, J.; Tian, Y.; Li, R.; Wang, B.; Lin, J.; Xu, A.; Huang, X.; et al. Differential Expression and Alternative Splicing of Transcripts Associated With Cisplatin-Induced Chemoresistance in Nasopharyngeal Carcinoma. Front. Genet. 2020, 11, 52. [Google Scholar] [CrossRef] [PubMed]
- Conrad, M. Transgenic mouse models for the vital selenoenzymes cytosolic thioredoxin reductase, mitochondrial thioredoxin reductase and glutathione peroxidase 4. BBA Gen. Subj. 2009, 1790, 1575–1585. [Google Scholar] [CrossRef] [PubMed]
- Michaelis, M.; Gralla, O.; Behrends, T.; Scharpf, M.; Endermann, T.; Rijntjes, E.; Pietschmann, N.; Hollenbach, B.; Schomburg, L. Selenoprotein P in seminal fluid is a novel biomarker of sperm quality. Biochem. Biophys. Res. Commun. 2014, 443, 905–910. [Google Scholar] [CrossRef]
- Yu, X.; Shi, W.; Cheng, L.; Wang, Y.; Chen, D.; Hu, X.; Xu, J.; Xu, L.; Wu, Y.; Qu, J.; et al. Identification of a rhodopsin gene mutation in a large family with autosomal dominant retinitis pigmentosa. Sci. Rep. 2016, 6, 19759. [Google Scholar] [CrossRef]






| Tissue | AS with Gene | Exon Skip** | Alternative 5′ Splice Site | Alternative 3′ Splice Site | Intron Retention |
|---|---|---|---|---|---|
| Brain_A | 147 | 113 | 60 | 52 | 58 |
| Breast | 126 | 92 | 80 | 71 | 63 |
| Testis_I | 150 | 80 | 71 | 70 | 99 |
| Lung | 202 | 94 | 95 | 75 | 225 |
| Placenta | 167 | 141 | 102 | 83 | 99 |
| Stomach | 89 | 48 | 56 | 48 | 34 |
| Gut_A | 144 | 76 | 111 | 54 | 86 |
| PBMC | 551 | 402 | 303 | 212 | 716 |
| Fat | 132 | 78 | 72 | 63 | 81 |
| Lymph | 317 | 133 | 497 | 116 | 479 |
| Prostate | 132 | 61 | 58 | 75 | 105 |
| Thymus | 238 | 215 | 194 | 118 | 224 |
| Heart_A | 116 | 89 | 55 | 72 | 56 |
| Brain_I | 192 | 101 | 54 | 81 | 116 |
| Gall | 175 | 78 | 70 | 64 | 146 |
| Nasopharynx | 157 | 84 | 87 | 82 | 92 |
| Retina | 252 | 192 | 113 | 104 | 228 |
| Thyroid | 316 | 114 | 82 | 79 | 350 |
| Liver_A | 108 | 97 | 69 | 66 | 45 |
| Gut_I | 150 | 77 | 69 | 77 | 107 |
| Oesophagus | 95 | 86 | 63 | 32 | 57 |
| Salivary | 143 | 67 | 128 | 48 | 117 |
| Urinary | 152 | 70 | 40 | 61 | 136 |
| Testis_A | 459 | 359 | 322 | 172 | 313 |
| Heart_I | 119 | 102 | 38 | 52 | 56 |
| Kidney | 140 | 156 | 113 | 88 | 78 |
| Ovary | 157 | 91 | 69 | 65 | 127 |
| Spinal | 170 | 84 | 84 | 80 | 102 |
| Uterus | 276 | 102 | 62 | 101 | 286 |
| Adrenal | 127 | 99 | 56 | 49 | 84 |
| Liver_I | 131 | 131 | 85 | 69 | 68 |
| Longissimus | 122 | 91 | 90 | 53 | 58 |
| Pancreas | 58 | 41 | 33 | 19 | 40 |
| Spleen | 128 | 117 | 56 | 64 | 75 |
| Position | Type | Canonical | Aberrant | Gene | Mutation | Detail |
|---|---|---|---|---|---|---|
| chr7: 22889860 | Novel | T28228 | T2132 | TMP-CH242-74M17.2 | Non-synonymous | ENSSSCT00000001330 |
| C188 | C17845 | ENSSSCG00000001229 | exon3:c.A395G:p.H132R | |||
| chr7: 22889835 | Novel | A27034 | A1996 | TMP-CH242-74M17.2 | Non-synonymous | ENSSSCT00000001329 |
| C0 | C2429 | ENSSSCG00000001229 | exon3:c.T370G | |||
| chr7: 22825399 | Novel | T2495 | T2956 | SLA-1 | Non-synonymous | ENSSSCT00000036412 |
| C0 | C2075 | ENSSSCG00000001231 | exon3:c.A404G:p.D135G | |||
| chr1: 236442561 | Novel | G2635 | G1216 | TLN1 | Non-synonymous | ENSSSCT00000005856 |
| T0 | T722 | ENSSSCG00000005317 | exon8:c.C745A:p.H249N | |||
| chr9: 67827403 | Novel | G1128 | G7 | C4BPA | Non-synonymous | ENSSSCT00000017061 |
| T593 | T7237 | ENSSSCG00000015663 | exon4:c.C605T:p.T202I | |||
| chr9: 67827383 | Novel | G1506 | G16 | C4BPA | Synonymous | ENSSSCT00000017061 |
| A520 | A8310 | ENSSSCG00000015663 | exon4:c.G585A:p.K195K | |||
| chr4: 90572872 | Semi-canonical | C17145 | C27 | TAGLN2 | Synonymous | ENSSSCT00000007008 |
| T7513 | T785 | ENSSSCG00000006395 | exon3:c.G246A:p.G82G | |||
| chr7: 1955628 | Semi-canonical | A6919 | A26 | TUBB2B | Synonymous | ENSSSCT00000001098 |
| G5 | G4623 | ENSSSCG00000001006 | exon1:c.T42C:p.N14N | |||
| chr6: 103414552 | Novel | A6090 | A79 | LOC733637 | Synonymous | ENSSSCT00000025362 |
| G14 | G325 | ENSSSCG00000003692 | exon4:c.T495C:p.F165F | |||
| chrX: 13203809 | Novel | T21856 | G0 | SYAP1 | Non-synonymous | ENSSSCT00000040956.2 |
| G0 | T4538 | ENSSSCG00000012148 | exon8:c.G970T:p.D295Y | |||
| chr7: 24913889 | Semi-canonical | C17224 | C14 | SLA-DRB1 | Synonymous | ENSSSCT00000001612 |
| T61 | T1190 | ENSSSCG00000001455 | exon1:c.G39A:p.A13A | |||
| chr6: 103414681 | Novel | A19996 | A114 | LOC733637 | Synonymous | ENSSSCT00000025362 |
| G481 | G332 | ENSSSCG00000003692 | exon4:c.T366C:p.G122G | |||
| chr7: 23636509 | Novel | C9015 | C3066 | SLA-7 | Synonymous | ENSSSCT00000035331 |
| T2779 | T2870 | ENSSSCG00000024161 | exon4:c.C852T:p.H284H |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Feng, W.; Zhao, P.; Zheng, X.; Hu, Z.; Liu, J. Profiling Novel Alternative Splicing within Multiple Tissues Provides Useful Insights into Porcine Genome Annotation. Genes 2020, 11, 1405. https://doi.org/10.3390/genes11121405
Feng W, Zhao P, Zheng X, Hu Z, Liu J. Profiling Novel Alternative Splicing within Multiple Tissues Provides Useful Insights into Porcine Genome Annotation. Genes. 2020; 11(12):1405. https://doi.org/10.3390/genes11121405
Chicago/Turabian StyleFeng, Wen, Pengju Zhao, **anrui Zheng, Zhengzheng Hu, and Jianfeng Liu. 2020. "Profiling Novel Alternative Splicing within Multiple Tissues Provides Useful Insights into Porcine Genome Annotation" Genes 11, no. 12: 1405. https://doi.org/10.3390/genes11121405