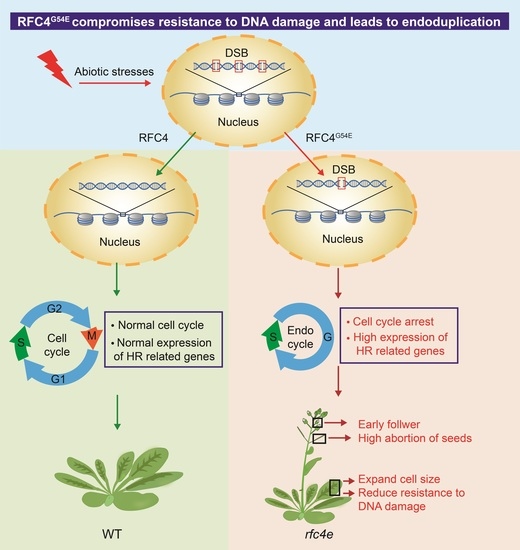
1. Introduction
The replication factor C (RFC) complex, which is an AAA+ATPase composed of one large subunit, known as RFC1, and four small subunits, RFC2/3/4/5, was first purified from the Hela cell extract of human cervical cancer and is essential for simian vacuolating virus 40 (SV40) DNA replication in vitro [
1,
2,
3,
4]. RFC subunits possess a cluster of conserved motifs that have been termed RFC boxes [
5]. The four small subunits, RFC2−5, and contain seven conserved RFC boxes, II−VIII, which mainly exist in the N-terminal region. RFC box III contains the most conserved motif: a phosphate-binding loop (P-loop or Walker A domain, GxxGxGK [S/T]) that is essential for the structure and function of RFC [
6,
7,
8]. While the conserved N-terminal structure is related to ATP binding/hydrolysis as well as binding to DNA [
4], the deletion mutations of RFC1/2/3/4/5 in humans [
9,
10],
Arabidopsis, and rice [
11] have shown that the C-terminal region is indispensable for RFC complex formation. RFC subunits assemble in a circular arrangement via tight interactions between their C-terminal domains; therefore, due to the indispensability of each subunit, the T−DNA insertion mutants of
AtRFC2−
5 are all lethal in
Arabidopsis [
11,
12].
Moreover, all five subunits are essential for maintaining the stability of the genome in eukaryotes. The
Scrfc1 (D513N) mutant exhibited a delay in the cell cycle, increased sensitivity to DNA-damaging agents and elongated telomeres in
Sacharomyces cerevisiae [
13]. Meanwhile, both
Scrfc2−1 (L304P
) and
Scrfc2−K71R have defects in the S-phase checkpoint, indicating that
ScRFC2 is important in both DNA replication and cell cycle checkpoints [
8,
14]. Temperature-sensitive
Scrfc5−1 (G43E) is sensitive to DNA-damaging agents, which can be suppressed by the overexpression of
Rad53, the essential protein kinase responsible for DNA damage, suggesting that RFC5 is part of a mechanism transducing the DNA damage signal and slowing the S-phase progression in response to DNA damage [
15,
16]. The
Sprfc3−1 (R216W) mutant showed aberrant mitosis with fragmented or unevenly separated chromosomes at a restricted temperature, as well as increased sensitivity to DNA-damaging agents and UV radiation [
17]. In addition, the mutation of
RFC4 in
Drosophila leads to the premature termination of protein translation, which causes striking defects in mitotic chromosome cohesion and condensation [
18].
In plants, genomic instability in RFC mutants may inhibit plant growth, trigger the DNA damage response (DDR), and lead to greatly increased frequencies of somatic recombination and heritable mutations, threatening cell survival and potentially leading to the transmission of deleterious mutations to the next generation [
19,
20,
21]. The DDR senses genomic damage and activates cell cycle checkpoints to promote DNA repair, and double-strand breaks (DSBs) in the DNA double helix are considered to be one of the major forms of DNA damage [
22]. Endoduplication is also one of the common responses to DNA damage in addition to cell cycle arrest and cell death. A lack of chromatin assembly factor 1 (CAF1) activity-induced DSB accelerates endoduplication in seedlings and leaves [
23]. Zeocin treatment in wild-type
Arabidopsis, which is known to produce DSBs, also shows increased DNA ploidy [
24].
Replication factor C subunit 1 (
RFC1) in
Arabidopsis thaliana was reported to play an important role in meiotic recombination and crossover formation, and in DNA double-strand break repair during meiosis [
25,
26,
27]. The
Atrfc3−1 (G84A) mutant exhibited hyper-sensitivity to salicylic acid and enhanced resistance to virulent oomycete
Hyaloperonospora arabidopsidis (
H. a.) Noco2, suggesting that
AtRFC3 negatively regulates systemic acquired resistance and has crucial functions in cell proliferation and DNA replication [
28,
29]. However, the molecular mechanisms of the other three subunits are still unclear due to the lack of viable mutants. In this study, to reveal the functional details of
AtRFC4 in higher plants, we successfully generated viable
rfc4−1/RFC4G54E (hereinafter referred to
rfc4e) mutants by site-directed mutagenesis in the Walker A domain of RFC4. The single amino acid substitution of 54th Gly to Glu in
AtRFC4 causes developmental defects and earlier flowering.
rfc4e mutants are also sensitive to DNA-damaging agents, leading to cell cycle arrest and endoduplication, indicating that
RFC4 is an important mediator of DNA damage repair in plant growth and resistance to abiotic stresses in
Arabidopsis.
2. Materials and Methods
2.1. Plant Materials and Growth Conditions
The AtRFC4 T−DNA line (Salk_049715) was acquired from the Arabidopsis Biological Resource Center. Plants were grown on ½ MS medium with 1% sucrose and 0.35% phytagel. Seeds were surface-sterilized for 2 min in bleach solution (added with 15% NaClO and 0.1% Tween 20), and then rinsed three times with sterilized water. The seeds were cold-treated for 3 days at 4 °C in the dark, and transferred to a growth chamber with long-day conditions of 16 h light and 8 h dark, at approximately 22 °C and 50% relative humidity, or directly planted on soil (nutrient soil: vermiculite: sand = 1:2:3); then, they were cultured in a growth room under 16 h light (22 °C)/8 h dark (22 °C) conditions.
2.2. Construction of Binary Vectors and Transformation
The site mutant variants of
RFC4(G54E/D) with native promotor were generated by PCR-mediated site-directed mutagenesis, as described in a published article [
30]. All primers used to generate
RFC4 site-specific mutants of
Arabidopsis are listed in
Table S4. The variants were sequenced and, via the ClonExpress II One Step Cloning Kit (Vazyme, C112-01, China), cloned into the
pCambia 1305::3FLAG vector and transformed into
rfc4−1 heterozygotes.
rfc4−1 plants were genotyped using the T
−DNA left border primer, LBa1 together with the
RFC4 gene-specific primer FP2 and the
RFC4 gene-specific primer pair FP1 and FP2. For the complementation test, the full-length genomic fragment of
AtRFC4, including its native promoter, was cloned into the
pCambia 1305::GFP plasmid vector and transformed to
rfc4e−3 plants.
All constructs were transformed into
Agrobacterium strain GV3101 using the electroporation method, and then transformed into plants using the floral-dip method, as described previously [
31]. Seeds harvested in bulk from each plant pot were sterilized and then screened on ½ MS medium supplemented with 50 μg·mL
−1 hygromycin. Selected homozygous mutants were used for phenoty** and transcription assays.
2.3. RNA Extract and RT-qPCR
For the expression analysis of genes, total RNA was isolated from whole seedlings using the Eastep™ Super Total RNA Extraction Kit (Promega, Madison, WI, USA). Reverse transcription was carried out using the GoScript™ Reverse Transcription System (Promega, Bei**g, China). The RT-qPCR assay was carried out using 2 × SYBR Green Premix Pro Taq HS Premix (AG11702, Accurate Biotechnology (Hunan) Co., Ltd., Changsha, China)) and a Step-One real-time fluorescence PCR instrument (Applied Biosystems, Bedford, MA, USA). The RT-qPCR reaction system contained 10 ng cDNA, 4 µM of each primer, 5 µL 2 × SYBR Green Premix Pro Taq HS Premix, 0.2 µL ROX reference dye and 3.4 µL RNAase-free water. The RT-qPCR programming was as follows: denaturation at 95 °C for 120 s, followed by 40 amplification cycles (95 °C for 20 s, 55 °C for 20 s and 72 °C for 30 s).
AtActin1 was used as an internal housekee** gene. Two or more independent biological replicates and three technical replicates of each sample were performed for quantitative PCR analysis. Gene-specific primers used in the experiments are listed in
Table S4.
2.4. Transcriptome Analysis
Total RNA was extracted by the mirVana™ miRNA Isolation Kit (Invitrogen, Waltham, MA, USA) from the 12
−d
−old Col
−0 and
rfc4e−3 mutant; plants were grown on a ½ MS plate in standard conditions. Library preparation and RNA sequencing were performed by OE Biotech Co., Ltd. (Shanghai, China). The paired-end RNA-seq sequencing library was sequenced with the Illumina Novaseq 6000 (2 × 150 bp read length) system. Three biological replicates per sample were analyzed. The clean reads were deposited into the NCBI Sequence Read Archive database (Accession Number: PRJNA820164).
p value < 0.05 and fold change ≥1.5 or fold change ≤0.67 were set as the thresholds for significantly differential expression. The GO enrichment analysis was based on the biological process functional categories of ShinyGo v0.75 software (
http://bioinformatics.Sdstate.edu/go/, date of access 1 March 2022) [
32].
2.5. True Leaf and Root Growth Inhibition Assays
In the true leaf inhibition assay, more than 100 seeds were sown and grown on a standard ½ MS plate, with or without DNA damage reagent, 15 μg·mL−1 Zeocin (Solarbio, Cas11006-33-0, Bei**g, China) or 80 ppm MMS (Sigma, Cas66-27-3, St. Louis, MO, USA). After 12 days, the number of true leaves was observed and counted for seedlings. For the root growth inhibition assay, seeds were sown and grown vertically on ½ MS plates, with or without DNA damage reagent, 3 μg·mL−1 Zeocin or 80 ppm MMS. After 7 days, their root growth was measured using ImageJ software.
2.6. Genotoxic Treatments
To investigate the expression of DDR genes after genotoxic treatment, 12−d−old seedlings grown under the growth conditions were transferred to and soaked in 100 μg·ml−1 Zeocin or 150 ppm MMS for 2 h. Seedlings were then harvested immediately after each stress treatment and subjected to gene expression analysis.
2.7. γ-H2AX Assays
For the γ-H2AX assays, total proteins were extracted with protein extraction buffer. The supernatant collected after centrifugation was transferred to a new tube, mixed with SDS loading buffer and boiled at 95 °C for 5 min. The protein samples were subjected to 12% SDS-PAGE gel electrophoresis, blotted and immunodetected with rabbit anti-γ-H2AX antibody (1 μg·mL−1, ab2893, Abcam, Cambridge, UK). AtGAPDH was used as an internal reference, detected using anti-GAPDH antibody (1:5000, 10494−1−AP, Proteintech Group, Inc Rosement, IL, USA). The band intensities on the immunoblots were determined by analyzing the exposed film with Quantity One professional grayscale analysis software.
2.8. Flow Cytometry
Approximately 0.5 cm
2 of the mature 1st to 2nd true rosette leaf of each plant (18
−d
−old) was chopped with a sharp razor blade in 400 µL of ice-cold CyStain UV Precise P nuclear lysis solution (Sysmex, Norderstedt, Germany), for 30
−60 s, to fully extract the complete cell nuclei. The nuclear suspension was filtered through a CellTrics 30 μm filter (Sysmex) directly into the sample tube, and 1600 µL of staining buffer (DAPI) was added. For the analysis of the nuclei, a Sysmex CyFlow
® Ploidy Analyser with UV excitation by a mercury arc lamp was used, and two or more samples were analyzed, with each sample containing at least 5000 nuclei. The data analysis was carried out with FCS Express version 3 software. The endoduplication index (EI) was calculated using the following equation: EI = (0×% 2C) + (1×% 4C) + (2×% 8C) + (3×% 16C) [
33].
2.9. Leaf Epidermal Cell Examination
The 1st to 2nd true leaf of each 18
−d
−old plant of the
rfc4e mutants and the wild-type Col
−0 was collected separately in a 15 mL tube and treated with a chloral hydrate:glycerol:water solution (8:1:2) to clear the cells [
34]. The epidermal cells on the abaxial leaf surfaces were photographed with a Zeiss Imager M2 microscope, and their leaf size was measured using ImageJ software.
4. Discussion
In the past few decades, many studies have provided details of the genes involved in the DNA replication and DNA damage repair process. However, due to the lethality of mutation, the four smaller subunits of the RFC complex have rarely been studied in higher plants. In this study, we generated viable mutants of
RFC4 with a single amino acid substitution by means of site-directed mutagenesis. Compared to Col
−0,
rfc4e mutants were smaller, flowered earlier, and produced a large number of aborted seeds, showing a DNA replication-defective phenotype (
Figure 1 and
Figure 2), similar to
rfc1−1 [
25] and
rfc3−1 [
29]. Interestingly,
rfc4e−1 showed a higher abortion rate than
rfc4e−2 and
rfc4e−3—this may be due to the relatively lower expression of
RFC4G54E in
rfc4e−1 (
Figure S2B). Consistent with this, cell division in embryos and endosperm was shown to be inhibited in
rfc4−1 heterozygotes, leading to the embryo lethality of the
rfc4−1 homozygotes in a former study [
12]. These results indicate that the single amino acid substitution in the conserved Walker A domain seriously affects the normal function of RFC4 and is not conducive to the adaptation of plants to the natural environment, but the abortion defect can be alleviated by an increase in
RFCG54E expression.
Transcriptome data analysis showed that the expression of DNA damage repair-related genes strongly increased in the
rfc4e−3 mutant, and immunoblotting experiments showed that there were more γ-H2AX signals in
rfc4e−1, rfc4e−2 and
rfc4e−3 mutants, indicating that
AtRFC4 mutation leads to an impaired ability to perform DSB repair. In plants, there are two main modules of DSB repair, HR and NHEJ. GO and KEGG aggregation analysis revealed that HR-related genes were highly expressed in
rfc4e mutants. Among these genes,
AtRAD51 is an essential recombinase for mitotic and meiotic HR repair [
37], while RAD54 [
38] is an essential cofactor that stimulates RAD51 activity, which has a significant effect on DNA damage repair in mitotic cells. In addition,
AtBRCA1 is also required for efficient DSB repair by homologous recombination in somatic cells [
36]. The significantly increased expression levels of these genes in
rfc4e mutants suggest that the HR repair pathway might be activated. However, there was no significant difference in the expression levels of NHEJ-related genes such as KU70/80, indicating that HR, rather than NHEJ, might be the main DSB repair mechanism in
rfc4e mutants. Consequently,
rfc4e mutants are supersensitive to DNA-damaging agents MMS and Zeocin, and they constitutively express higher levels of DDR-related genes, such as
GR1,
BRCA1,
RAD51,
PARP1, PARP2 and
RPA1E. Mutations in core replication machinery proteins (such as Pol δ [
47]) and replication proteins RFC1 [
25] and RAD17 [
45] are known to increase somatic homologous recombination (SHR), and they are sensitive to DNA-damaging agents. Moreover, the expression of DDR genes in
rfc3−1 mutants shows a similar expression pattern to that observed in
rfc4e mutants, further suggesting that the mutation of these core DNA replication factors compromises the capacity for DSB repair during DNA replication and DNA damage in
rfc4e and
rfc3−1 mutants.
When DNA damage occurs, plant cells have the choice of either delaying cell division to repair the damage or inducing cell death. A likely benefit of the first choice is that entering into endoduplication prevents DNA-damaged cells from proliferating and also from dying. In
rfc4e mutants, we observed lower numbers of epidermal cells with increased DNA ploidy. Several genes that have been identified as critical for the inward replication of cyclins and cyclin-dependent kinase (CDK) complexes are all upregulated in
rfc4e mutants. The
Arabidopsis mitotic cyclin
CYCB1;1 is expressed during the G2/M transition, and its upregulation suggests a G2 arrest [
48]. Members of the SIAMESE/SIAMESE-RELATED (SIM/SMR) class of cyclin-dependent kinase inhibitors, including SMR4, SMR5 and SMR7, activate cell cycle checkpoints in response to DNA damage, thereby inhibiting mitosis and promoting endoduplication [
49]. The high expression of cell cycle inhibitory protein kinase WEE1 can promote nuclear replication, and its expression in tomato fruit tissue is positively correlated with DNA fold [
50]. In
Drosophila, the mitotic cells of
I(3)Rfc4e18 and
I(3)Rfc4e20 lines exhibited prematurely condensed chromosomes or chromosomal bridges/breaks [
18]. Meanwhile, mutations of
Scrfc2,
Scrfc5,
Sprfc2 and
Sprfc3 caused replication defects and chromosomal abnormalities when entering the mitotic stage [
14,
16,
17,
51,
52]. Based on this evidence, it is plausible to postulate that aberrant mitosis takes place in
rfc4e mutants. Combined with the increased DSB, embryonic abortion, high expression of DDR genes and endoduplication in
rfc4e mutants, our data suggest that
AtRFC4 plays an important role in maintaining normal DNA replication and damage repair, and a single amino acid substitution will lead to genomic instability, an impaired repair ability and inward replication transformation of the cell cycle, which ultimately compromises cell proliferation and resistance to genotoxic stresses.
Although there are many methods to study gene function, there are still some genes, which, due to their importance and irreplaceability, are unable to obtain corresponding mutants using EMS mutagenesis or T-DNA insertion methods. The acquisition of RFC4 single amino acid substitution mutants not only enriches the biological function of RFC4 in plants, but also provides a means to study similar housekee** genes. Above all, the acquisition of mutants is conducive to studying the comprehensive effects of these genes in plants.