1. Introduction
R-loops form when a complementary RNA molecule invades the DNA double helix and hybridizes with one strand through Watson–Crick pairing, leaving the second DNA strand single-stranded. It is believed that most R-loops form in a co-transcriptional manner, when the nascent RNA invades its template in the wake of the RNA polymerase (reviewed in [
1,
2,
3]). In eukaryotes, R-loops can form both at protein-coding and non-coding transcription units, whatever the RNA Polymerase involved and it is estimated that 5% of the human genome has the potential to form detectable R-loops [
4]. R-loop biology is the focus of growing scrutiny, as increasing evidence suggest that the defective control of R-loop formation is associated with a number of human diseases [
5]. Many questions remain however regarding the direct consequences of R-loop formation on the surrounding chromatin.
Current data demonstrate that specific features facilitate R-loop formation. These include negative topological stress in the DNA, a strong bias for Guanine residues (G-skew) or the presence of homopolymeric Adenine tracts in the RNA sequence, and high rates of transcription (reviewed in [
2]). Some of these features are inter-dependent, as high rates of transcription will lead to greater negative topological stress, and, incidentally, will also increase the likelihood of forming and hence detecting R-loops. Nevertheless, a consensus is emerging that the beginning of G-skewed genes represents strong R-loop forming regions [
4,
6,
7,
8], although it could be that those particular R-loops are thermodynamically more stable because of the G-skew, and hence easier to detect. Using some map** methods at least, R-loops have also been shown to form in gene bodies and in terminator regions [
4], although the presence of R-loops in gene terminators was not significantly detected in
Arabidopsis thaliana [
8]. Recent observations have suggested that promoter R-loops tend to form over DNA sequences where RNA Polymerase II is frequently in pause [
6,
9,
10] and that have the potential to form G-quadruplex structures [
6]. It is possible that R-loops, by displacing G-rich single-stranded DNA (ssDNA), facilitate the formation of intra-molecular G-quadruplexes, which in turn, could stabilize R-loops. Taken together, these observations clearly show that there are strong sequence determinants to R-loop formation.
In eukaryotes, chromatin structure (i.e., the position and dynamic of nucleosomes) also modulates R-loop formation [
11,
12,
13], presumably because the presence of nucleosomes counter-acts the propensity of the double-helix to open and be invaded by the nascent RNA. The folding of the nascent RNA and its coating by RNA-binding proteins such as the spliceosome is also believed to strongly antagonize R-loop formation, most likely by physically preventing the nascent RNA from threading back into its DNA template [
14,
15]. Consistent with this, the presence of introns was shown to limit R-loop formation [
15] and conversely, R-loop formation was shown to increase significantly in many mutants of the RNA processing machinery [
16,
17]. Note however that, somewhat in apparent contradiction with this simple model, there is evidence that the homologous recombination (HR) machinery is essential for R-loop formation in many RNA processing mutants in budding yeast, suggesting that R-loop formation is an active process that requires the Rad51 recombinase [
18]. The significance of this observation is not yet fully understood. More importantly, recent data suggest that DNA replication has a strong influence on R-loop formation [
19,
20]: head-on conflicts between the transcription and replication would facilitate R-loop formation, possibly because topological constraints associated with such conflicts induce more negative topological stress over R-loop forming genes, whilst co-directional conflicts would reduce R-loop formation, possibly because replisome-associated helicases could disassemble R-loops. In addition, recent data suggest that in human cells at least, R-loop formation in mitosis is largely restricted to centromeres [
21]. Taken together, these observations strongly suggest that the cell cycle stage greatly influences the abundance and localization of R-loops. To sum up, although our understanding of the features that modulate R-loop formation has increased considerably in recent years, in-depth and consensual mechanistic details of R-loop formation at the molecular level are still missing.
R-loops have been previously proposed to act as double-edged swords in the genome [
1], because they can be both physiological intermediates in some key processes and pathological structures. For example, R-loops have been shown to participate in class-switch recombination at immunoglobulin genes [
22,
23], in the initiation of mitochondrial replication [
24], in the hypo-methylation of promoters [
25,
26] or in transcription termination [
27,
28,
29]. Recently, R-loops have also been proposed to form on either side of double-strand breaks [
30]. Conversely, R-loops are mutagenic [
31], they stimulate transcription-associated recombination (TAR) [
32], and they are associated with DNA damage in a replication-dependent manner [
33]. In particular, R-loops are hugely detrimental in head-on collisions between transcription and replication, where they lead to severe genome instability [
19,
20]. What features determine when R-loops become pathological structures remains poorly understood, but there is general agreement that R-loop stability and the chromatin context in which they form are likely to be key contributing factors. It is also possible that there are in fact different types of R-loops in the cell, depending, for example, on their size or their sequence, and that those different R-loops have different levels of toxicity depending on their architecture or the set of proteins that recognize them. Consistent with the idea that there could be different types of R-loops, R-loops that form in the absence of the THO complex in
Saccharomyces cerevisiae display much higher levels of TAR than those that form when both ribonuclease (RNase) H1 and H2 are missing [
32].
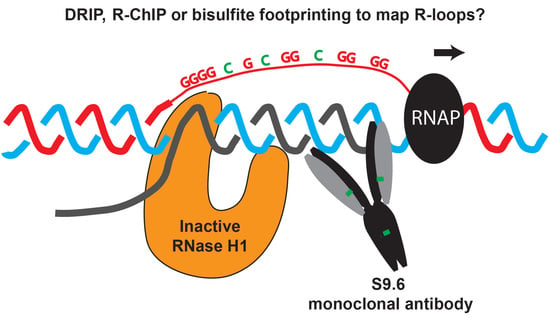
To address further the possible functions of R-loops, reproducible, accurate, and quantitative R-loop map** techniques are needed. The most widely used of these map** techniques is DNA:RNA immunoprecipitation (DRIP) [
25] that relies on the use of the S9.6 antibody, which recognizes DNA:RNA hybrids with great affinity [
34]. Recently, several studies have questioned the robustness of this approach [
6,
7,
35] and an alternative strategy to map R-loops has been developed [
6]. In addition, the understanding of R-loop functions necessitates robust and specific approaches to modulate R-loop formation. For this, the available toolkit remains very limited and most studies rely on the nuclear over-expression of RNase H, an enzyme that destroys R-loops without specificity, whether they are physiological or pathological.
Here, I aim to conduct an impartial assessment of the strengths and weaknesses of the different methods to map and evaluate the functions of R-loops. Based on these observations, I propose guidelines for best practices when working with R-loops.
4. RNase H1-Based Methods to Map R-Loops
RNase H1 is a highly conserved enzyme that recognizes DNA:RNA hybrids and cleaves their RNA moiety. In human cells, the low abundant RNase H1 is particularly enriched in mitochondria and in the nucleolus in a transcription-dependent manner [
42]. Its role in the nucleus is not yet very well characterized however. As it is an endogenous activity that evolved to recognize and to disassemble DNA:RNA hybrids, RNase H1 has been used as a tool to map R-loops. In particular, several studies have used a mutant of RNase H1 that could recognize but not process DNA:RNA hybrids [
6,
25,
43,
53]. The earliest strategy was to express this catalytically inactive human RNase H1 (hRNase H1-D145N) in bacteria and to couple it to amylose beads. R-loops were then purified by affinity from sheared genomic DNA. This strategy, called DNA:RNA in vitro enrichment (DRIVE-seq), identified fewer R-loops than DRIP-seq [
25,
53]. To improve the sensitivity of this strategy, we made the same mutation in the endogenous RNase H1 enzyme in fission yeast (Rnh1-D129N), and we used chromatin immunoprecipitation (ChIP) of Rnh1-D129N to demonstrate the formation of unstable R-loops at tRNA genes [
43]. The enrichment of Rnh1D129N at tRNA genes was fully sensitive to the strong in vivo expression of catalytically-active RNase H1 from
Escherichia coli (RnhA), validating the use of Rnh1D129N as an R-loop reporter at these sites [
43]. A similar approach was recently implemented in human cells using the nuclear over-expression of the catalytically inactive RNase H1-D210N and was renamed R-ChIP [
6]. Importantly, the sequencing of R-ChIP reactions in this latest study was strand-specific and was controlled using a mutant of RNase H1 that cannot recognize DNA:RNA hybrids [
6].
As with DRIVE-seq, R-ChIP identified fewer R-loop forming regions than the S9.6-based DRIP-seq or DRIPc-seq approaches [
6]. Importantly, R-ChIP identified R-loops mostly at promoters, but not at terminator regions [
6]. In addition, R-ChIP identified smaller R-loop forming regions than DRIPc-seq [
6]. The size of R-loops identified by R-ChIP was very similar to the archetypical size of R-loops mapped in vitro at nucleotide resolution using non-denaturing bisulfite footprinting. Moreover, the R-loop forming regions identified by R-ChIP contained sequence motifs, such as clusters of Gs in the non-template strand that were previously identified in vitro as potential triggers for R-loop formation [
54]. Finally, in both fission yeast and human cells, R-ChIP identified tRNA genes as hotspots of R-loop formation in otherwise wild-type cells [
6,
43], whilst tRNAs were identified as R-loop forming regions by DRIP in
A. thaliana [
8], but not in human cells [
4,
25]. Taken together, these observations led Chen et al. [
6] to conclude that R-ChIP is better at identifying genuine R-loops than S9.6-based methods.
There are however significant down sides to R-ChIP. Although it is easy to implement in yeast [
43], it is harder to implement in vertebrate cells, because it requires the stable expression of a mutant enzyme. As shown recently [
21], the over-expression of a catalytically-inactive RNase H1 enzyme presents the risk of interfering with the dynamics of R-loops in vivo, when there is good evidence that R-loop dynamics is likely to be critical for gene expression and genome stability. To implement R-ChIP in mammalian cells, it is therefore important that the catalytically-inactive RNase H1 is expressed at the right level: too much expression and there is a risk of dominant-negative effects; too little expression and there is a risk that the endogenous, catalytically-active enzyme could interfere with the binding of the catalytically-inactive mutant and the efficiency of R-ChIP. More importantly even, there is a significant risk that R-ChIP is only going to map the R-loops that are recognized by RNase H1. As an increasing number of proteins have been postulated to recognize and disassemble R-loops in vivo (see for example Senataxin [
55], FANCM [
56], BLM [
57], DDX19 [
58], MTR4 [
31] among others), it is conceivable that there might be different types of R-loops that could be recognized by different types of proteins. This might be why R-ChIP did not identify terminator regions as R-loop hotspots in human cells [
6], where R-loops might be recognized and disassembled by DNA&RNA helicases, such as Senataxin and not by RNase H1 [
27]. This could also explain why R-ChIP and DRIVE-seq identified fewer R-loop forming regions than DRIP-seq or DRIPc-seq.
We argued previously that R-ChIP and DRIP are complementary approaches because their use in parallel could give information about the stability of R-loops at specific loci [
43]. For example in fission yeast, RNase H1 is most abundant at RNAPIII-transcribed genes, suggesting that R-loops are constantly formed and detected there [
43]. The R-ChIP signal is therefore very strong at RNAPIII-transcribed genes and this was also true in human cells [
6]. On the contrary, DRIP-qPCR and DRIPc-seq only gave significant signals at RNAPIII-transcribed genes in fission yeast in the absence of RNase H1 and RNase H2 [
38,
43], thus confirming that RNAPIII-transcribed genes produce R-loops that are efficiently degraded by RNase H enzymes. To summarize these observations, it is conceivable that DRIP is better suited at detecting long-lived R-loops, whilst R-ChIP could be better at detecting highly dynamic R-loops that are processed by RNase H1.
5. RNase H Over-Expression as a Tool to Probe R-Loop Functions
As discussed above, the direct consequences of R-loop formation on the surrounding chromatin are still largely unclear. An in-depth understanding of R-loop contribution to gene expression and genome stability necessitates functional assays where R-loop formation could be specifically modulated. Importantly, to secure an unequivocal interpretation of the data, R-loop formation should be modulated without interfering with transcript synthesis or integrity.
The most common functional assay to probe R-loop functions relies on the long-term modulation of R-loop levels by affecting RNase H activity in vivo: RNase H-sensitive R-loops are classically stabilized by deleting or down-regulating RNase H enzymes and in most model systems, RNase H-sensitive R-loops can be de-stabilized by artificially increasing the concentration of RNase H enzymes in the nucleus. When increasing (or decreasing) RNase H activity in the nucleus alters a phenotype of interest, it is concluded that R-loops contribute to this phenotype. Note however that this approach does not demonstrate that the contribution of R-loops to the phenotype of interest is direct or indirect.
The most widely used strategy is to over-express RNase H. Although being widely used, this strategy presents significant disadvantages. Its biggest limitation is that the amount of R-loops is reduced genome-wide and without specificity, meaning that both the physiological and the pathological RNase H-sensitive R-loops are affected. In addition, although this has never been shown, there is at least a theoretical risk that a strong concentration of RNase H in the nucleus might also interfere with the steady-state of other DNA:RNA hybrids, such as the primers of Okazaki fragments or the DNA:RNA hybrids that were recently detected at double-strand breaks during repair [
30,
59]. This could be why RNase H1 over-expression in human cells was associated with persistent DNA damage [
42]. Finally, as discussed above, the possibility that there are RNase H-resistant R-loops in vivo that would resist such treatment has not yet been excluded.
Importantly, the fact that RNase H over-expression has an indiscriminate effect on DNA:RNA hybrids opens the possibility that it will have a significant impact on the transcriptome and the proteome, and that those changes could indirectly contribute to alter the phenotype of interest. If true, this would seriously complicate the interpretation of such experiments. For example, it was shown that the over-expression of RNase H1 in human cells significantly affects the protein levels of Top1 and other DNA repair proteins [
42]. In addition, we recently demonstrated that the strong expression of
E. coli RnhA in fission yeast had a significant impact on the transcriptome and affected the steady-state levels of many RNAs that did not form R-loops, according to our DRIPc-seq maps [
38]. These observations confirmed that long-term manipulation of RNase H activity imparts significant and indirect changes to the transcriptome and proteome. In addition, the transcriptome and proteome alterations that are associated with RNase H over-expression are likely to differ in different genetic backgrounds. In particular, the modifications to the transcriptome imparted by RNase H over-expression could combine with the existing alterations to the transcriptome in some mutant backgrounds. Consistent with this, our unpublished results indicate that the strong expression of RnhA in fission yeast differently alters the transcriptome in different mutant backgrounds. Therefore, the fact that a phenotype of interest in a mutant background is sensitive to RNase H over-expression does not necessarily mean that R-loops contribute to this phenotype directly. Nevertheless, if this strategy were to be implemented to probe R-loop function because of its relative ease, it should probably be backed up by the demonstration that the over-expression of another R-loop removing enzyme also impacts the phenotype of interest in a similar way. Alternatively, ectopic expression of AID, a cytidine deaminase that targets cytosine residues in ssDNA, was shown previously to enhance R-loop dependent recombination and mutagenesis in yeast [
13,
60], and could be used to probe R-loop functions at the genome-wide level. Note however that AID could also target ssDNA present in other forms of non-B DNA [
61], which could complicate the interpretation of such experiments.
However, if the consequences of R-loop formation on gene expression and genome stability are highly dependent on the chromatin context in which they form, locus-specific ways of manipulating R-loops must be developed. A corollary to what was discussed above is that locus-specific assays should be preferred to genome-wide approaches anyway because they are probably at lower risk of imparting indirect effects. To modulate R-loop formation locally, promoter-inactivating mutations [
62] or ribozyme-induced transcript cleavage [
32] have been used previously. The down side of these approaches is that they affect R-loop formation by interfering with the synthesis or the integrity of the transcript itself. Unless one provides strong arguments establishing that the sole purpose of transcription at the locus of interest is to form R-loops, the interpretation of such experiments can be complicated because they remove both the R-loop and the transcript. Ideally, one should aim to modulate the R-loop formation without affecting the synthesis of the transcript. To increase the concentration of RNase H (or other R-loop removing enzymes) locally using chromosome-targeting approaches could be an alternative strategy. A down side to this approach is that it is possible that the chromosome-targeting mechanism would interfere with the activity, the dynamics or efficiency of RNase H, which would result in only an incomplete reduction of R-loop levels. Another possibility could be to use genome editing to alter the sequence determinants leading to R-loop formation in a sequence of interest, without interfering with the synthesis of the transcript. Although this should be in theory the best approach, it also presents down sides: it is cumbersome to implement because the sequence determinants would have to be identified and validated in vitro before being mutated in vivo; in addition, it would be difficult to alter such sequence determinants in protein-coding sequences without altering the protein sequence itself, because some amino-acids, such as Glycine, are exclusively encoded by G-rich codons. Depending on the phenotype that is investigated, this could complicate the interpretation of the results. This approach might therefore be better implemented to understand the role of R-loop formation within non-coding transcription units.
To conclude, to go further forward in the in-depth characterization of R-loop contributions to gene expression and genome stability, there are some serious technical hurdles: RNase H over-expression, which is the strategy that is easiest to implement, is subject to potential indirect effects, whilst locus-specific strategies that would be better adapted at evaluating the direct consequences of R-loop formation on the surrounding chromatin are both risky and difficult to implement. Consensual proof-of-concept experiments are required to move the field forward.